Benjamin Midtvedt, Jesús Pineda, Fredrik Skärberg, Erik Olsén, Harshith Bachimanchi, Emelie Wesén, Elin K. Esbjörner, Erik Selander, Fredrik Höök, Daniel Midtvedt, Giovanni Volpe
Nature Communications 13, 7492 (2022)
arXiv: 2202.13546
doi: 10.1038/s41467-022-35004-y
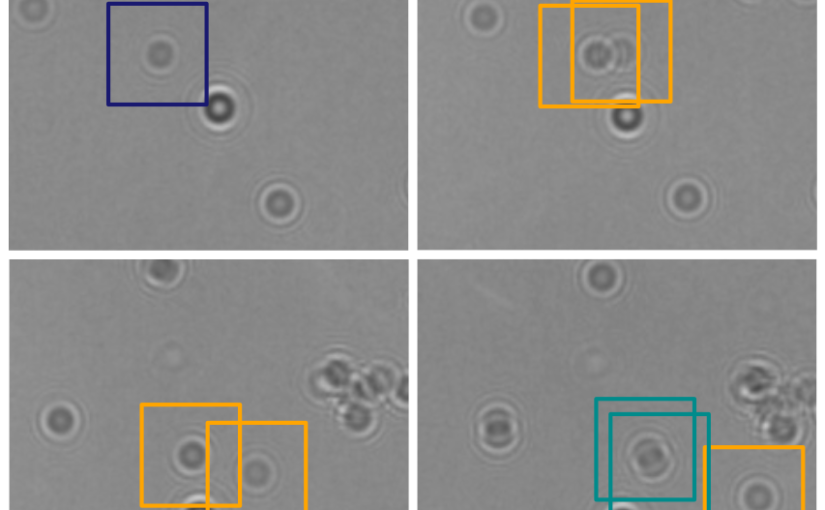
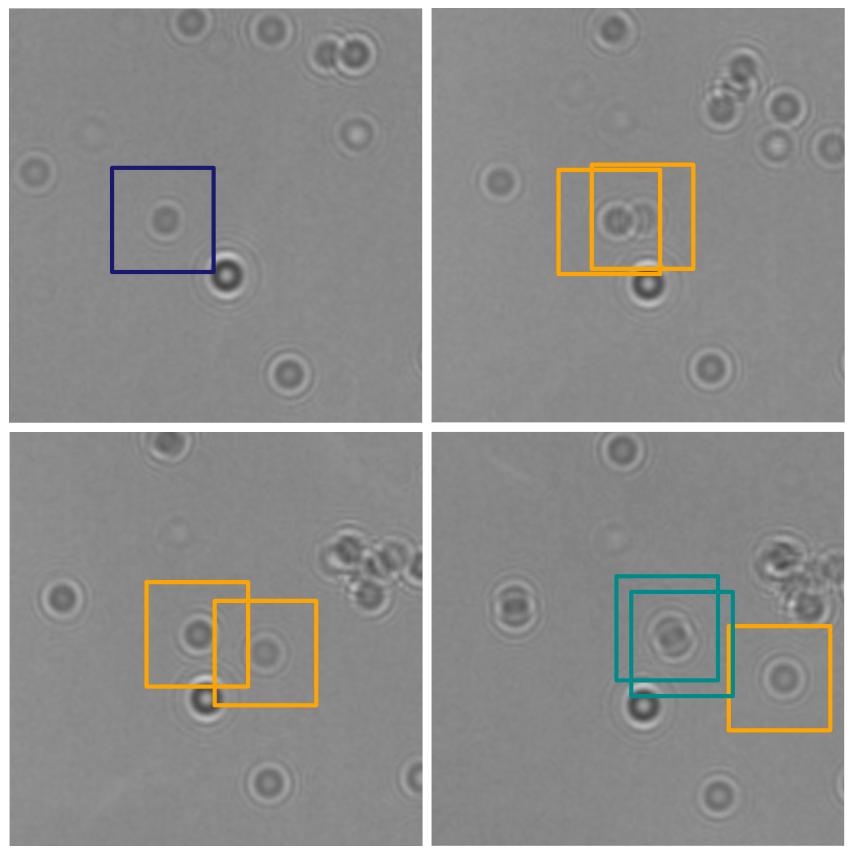
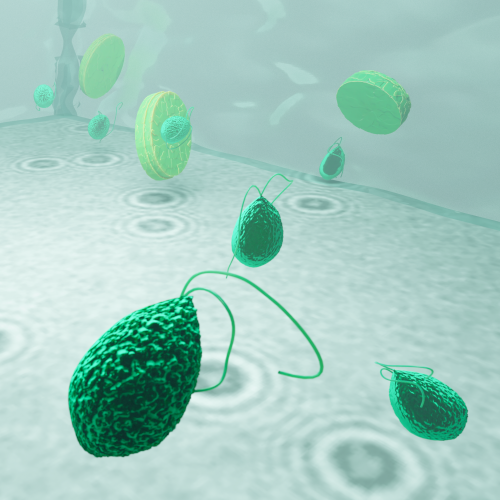
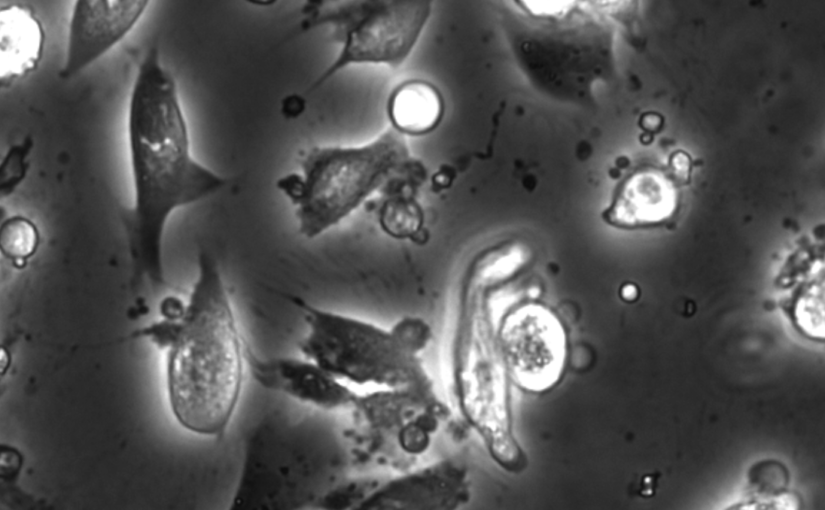
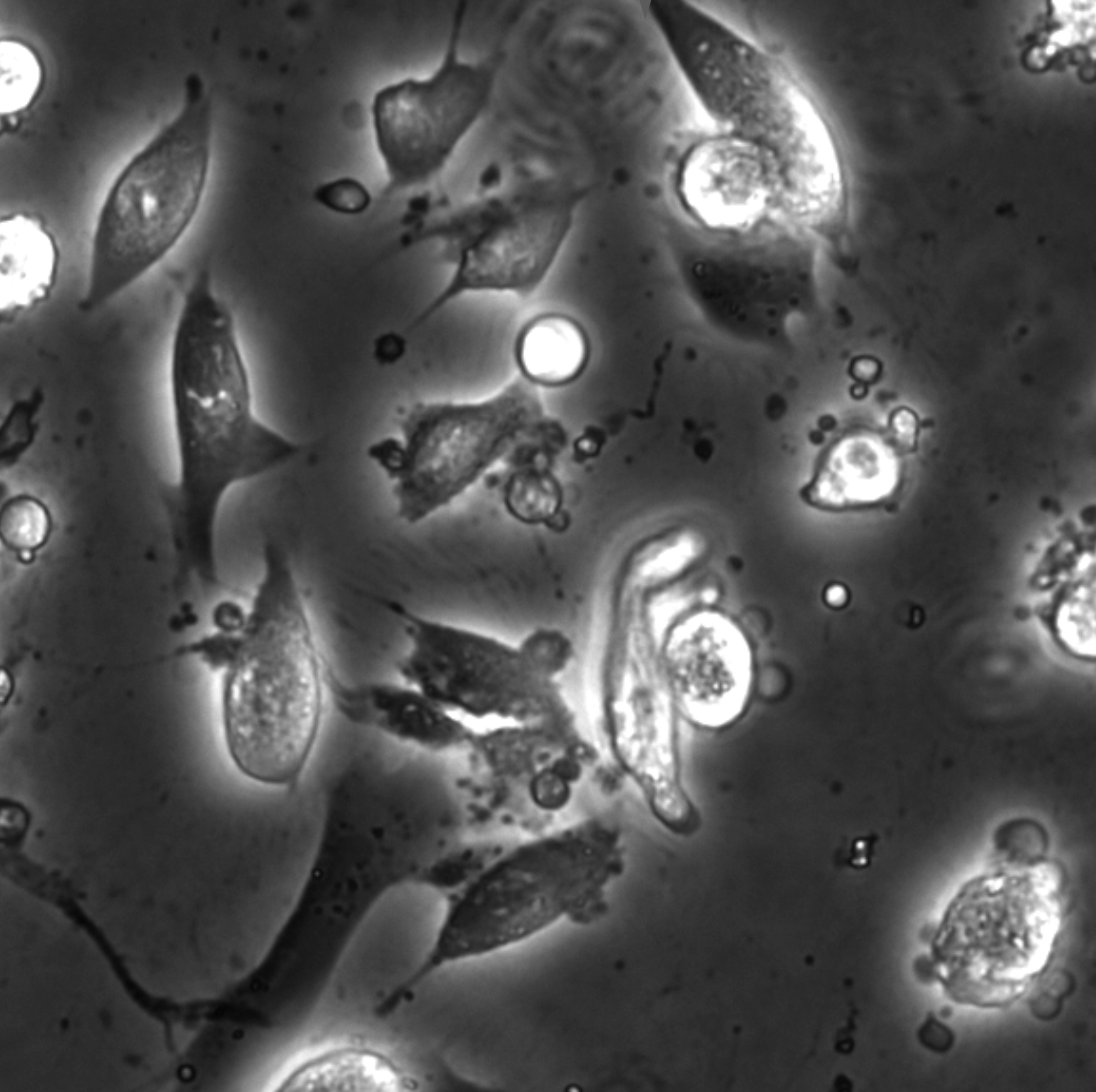
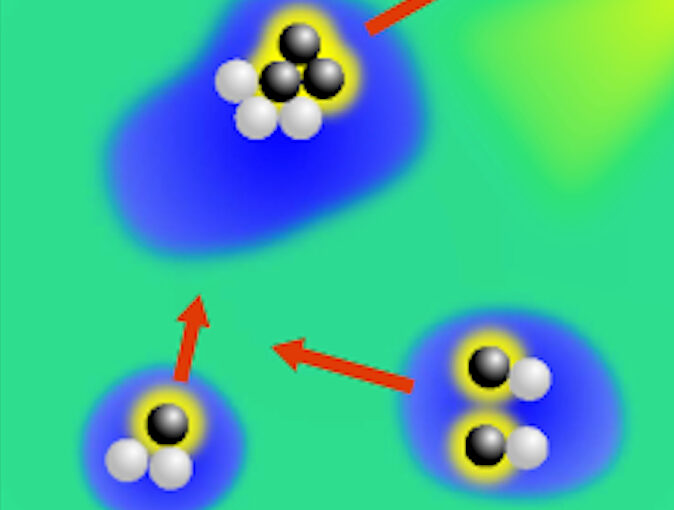
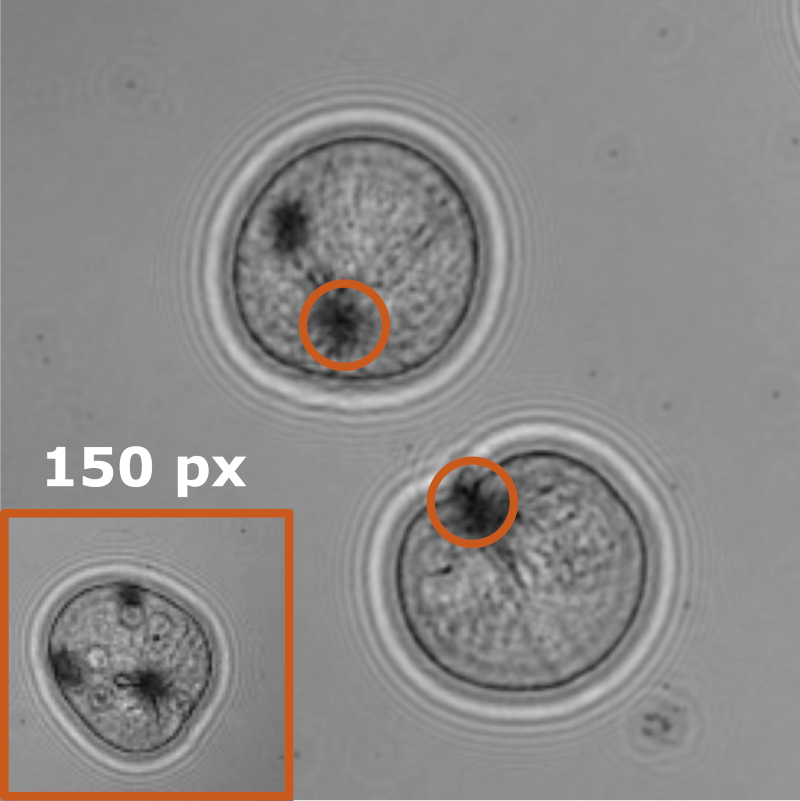
Object detection is a fundamental task in digital microscopy, where machine learning has made great strides in overcoming the limitations of classical approaches. The training of state-of-the-art machine-learning methods almost universally relies on vast amounts of labeled experimental data or the ability to numerically simulate realistic datasets. However, experimental data are often challenging to label and cannot be easily reproduced numerically. Here, we propose a deep-learning method, named LodeSTAR (Localization and detection from Symmetries, Translations And Rotations), that learns to detect microscopic objects with sub-pixel accuracy from a single unlabeled experimental image by exploiting the inherent roto-translational symmetries of this task. We demonstrate that LodeSTAR outperforms traditional methods in terms of accuracy, also when analyzing challenging experimental data containing densely packed cells or noisy backgrounds. Furthermore, by exploiting additional symmetries we show that LodeSTAR can measure other properties, e.g., vertical position and polarizability in holographic microscopy.