Viktor Holubec, Alexander Fischer, Giovanni Volpe, Frank Cichos
Physical Review Research 8, L022017 (2026)
arXiv: 2505.11042
doi: 10.1103/xn9x-ppjx
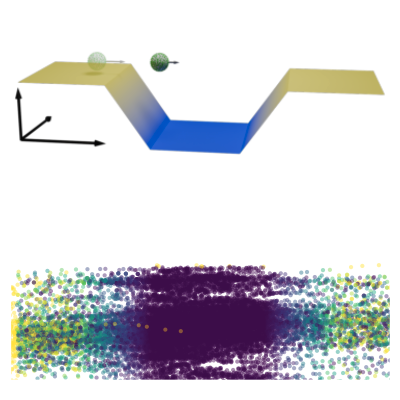
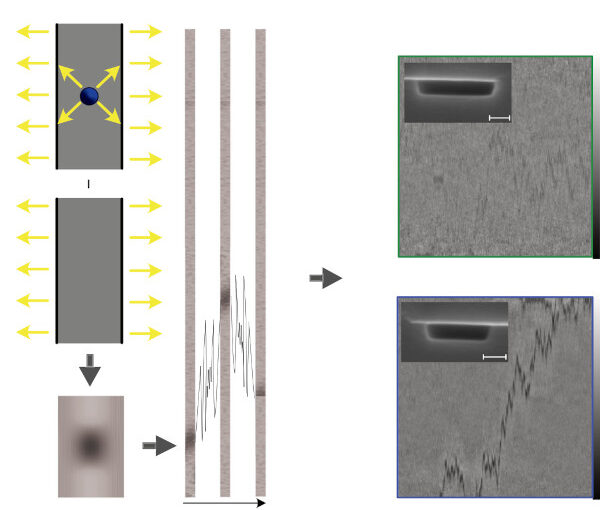
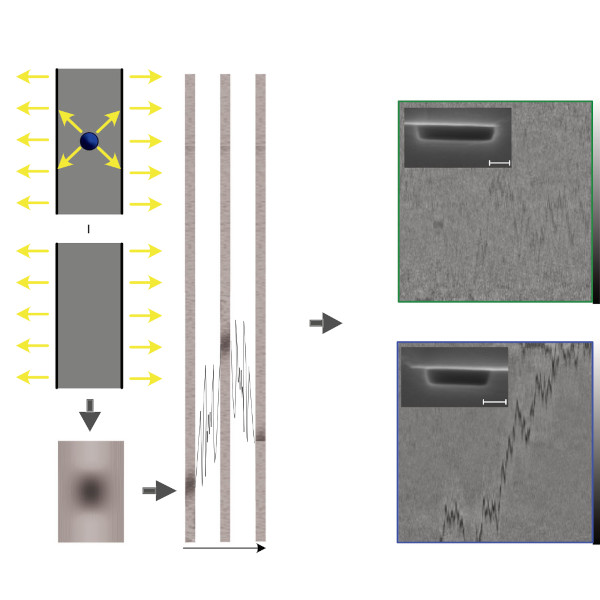
Active systems in nature and synthetic environments commonly exhibit spatially heterogeneous activity patterns and time-delayed responses from internal feedback mechanisms, exemplified by bacterial chemotaxis. We study an idealized active gas where particles modulate their self-propulsion based on local environmental conditions with such delays. Through integrated theoretical, computational, and experimental approaches, we demonstrate that steady-state density distributions and collective polarization exhibit characteristic peaks and valleys as functions of response delay time. We find that delays can amplify polarization by nearly an order of magnitude and trigger complete polarization reversal when particle displacement during the delay period surpasses the persistence length. Multiparticle simulations incorporating interparticle interactions validate that these phenomena remain robust in sufficiently dilute collective systems. Since density and polarization determine the current in active matter, our findings show that temporally programming the delay time allows control over both static and dynamic states in active systems, with implications for biological microswimmers and engineered microrobots.