The defense took place in PJ, Institutionen för fysik, Origovägen 6b, Göteborg, at 13:00.
Title: Deep Learning Enhanced Optical Methods for Single-Plankton Studies
Abstract: Among Earth’s earliest life forms, cyanobacteria reshaped the planet by oxygenating the atmosphere during the Great Oxidation Event 2.4 billion years ago. This process, which led to ozone formation and UV protection, paved the way for more complex photosynthetic organisms—phytoplankton, the eukaryotic descendants of cyanobacteria. Today, phytoplankton drive the global carbon cycle, producing 50–80% of Earth’s oxygen and fueling the marine food web. Microzooplankton consume nearly two-thirds of the organic carbon generated, yet despite their ecological significance, tracking biomass flow at the single-cell level remains a major challenge.
This thesis presents novel methodologies that integrate advanced optical techniques, deep learning, and simulated datasets to analyze microplankton dynamics with unprecedented resolution.
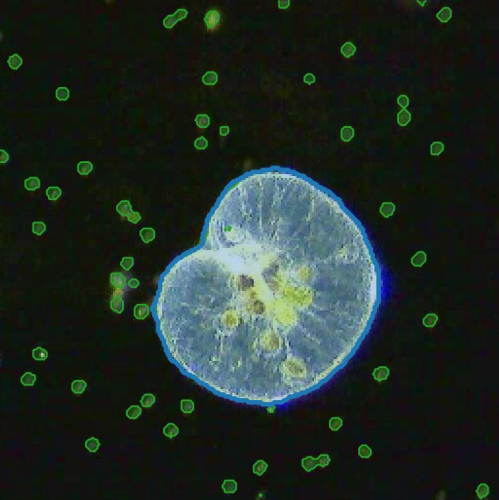
A key contribution is a deep-learning-enhanced holographic microscopy approach that quantifies microplankton biomass at the single-cell level while simultaneously capturing their three-dimensional swimming behavior. This method overcomes computational bottlenecks in traditional holography, enabling high-throughput analysis across diverse species and size ranges. Expanding on this, I demonstrate its application in mixed-species experiments to examine feeding interactions between phytoplankton and microzooplankton, capturing biomass transfer and behavioral shifts during predation.
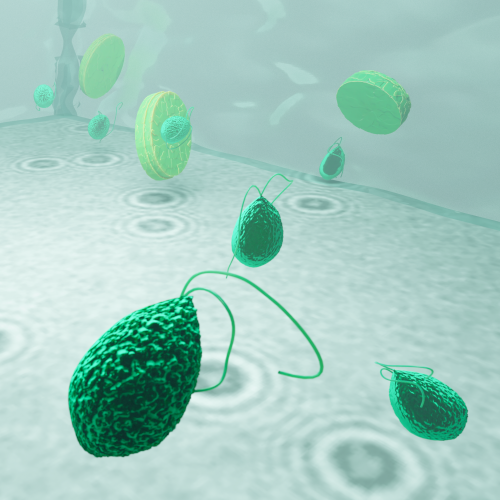
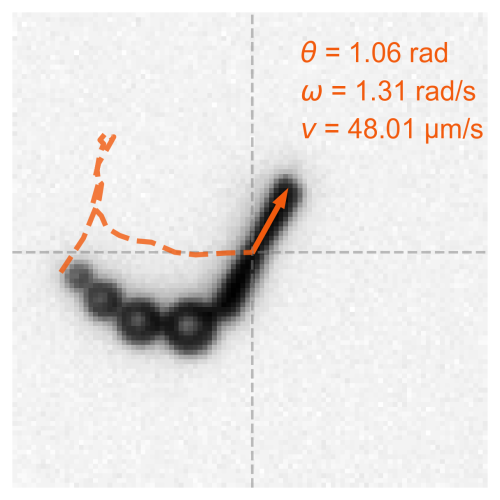
Beyond direct imaging, this thesis leverages synthetic data to advance microscopy-based research. Neural networks trained on simulated microscopy datasets are used to detect, segment, and classify plankton species while reconstructing motion dynamics. To showcase the versatility of this approach, I present its application in a non-biological setting—detecting bubble-propelled artificial micromotors within complex experimental backgrounds. In addition to object detection, these methods also enable motion characterization of microscopic entities. To demonstrate this, I introduce synthetic microscopy videos that model microscopic organisms undergoing various anomalous diffusion behaviors. This framework is then used to develop a method that extracts motion characteristics without explicit trajectory linking, broadening its applications beyond plankton ecology.
Finally, I investigate how zooplankton—key players in the marine food web—respond to ocean wave-induced light patterns using an LED matrix. The results suggest that zooplankton use steady light sources, such as celestial objects, to ascend more rapidly during favorable low-turbulent conditions, offering new insights into their migratory strategies. Collectively, this thesis bridges marine ecology, microscopy, artificial intelligence, and biophysics to provide new tools for exploring the unseen dynamics that shape our planet.
Thesis: https://hdl.handle.net/2077/84857
Supervisor: Giovanni Volpe
Examiner: Raimund Feifel
Opponent: Anupam Sengupta
Committee: Elisa Berdalet, Maria Guix Noguera, Josefin Titelman
Alternate board member: Paolo Vinai