Carlos Bustamante, Christian Kaiser, Erik Lindahl, Robert Sosa, Giovanni Volpe
arXiv: 2510.27074
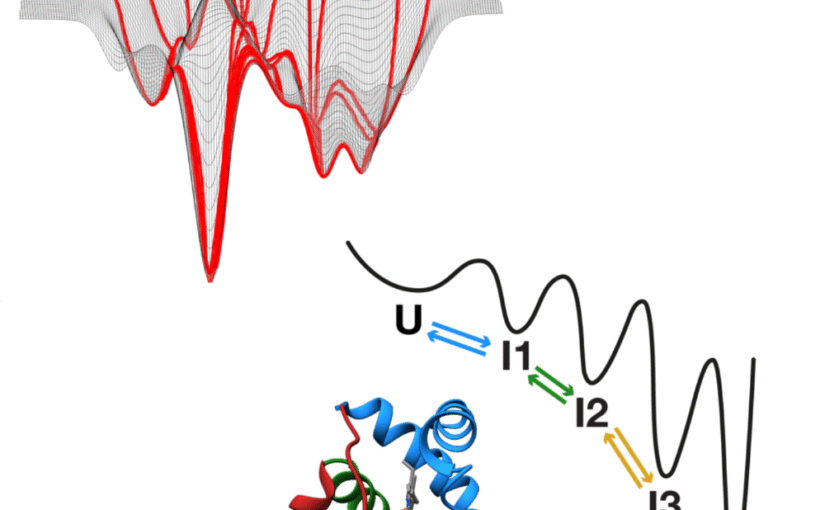
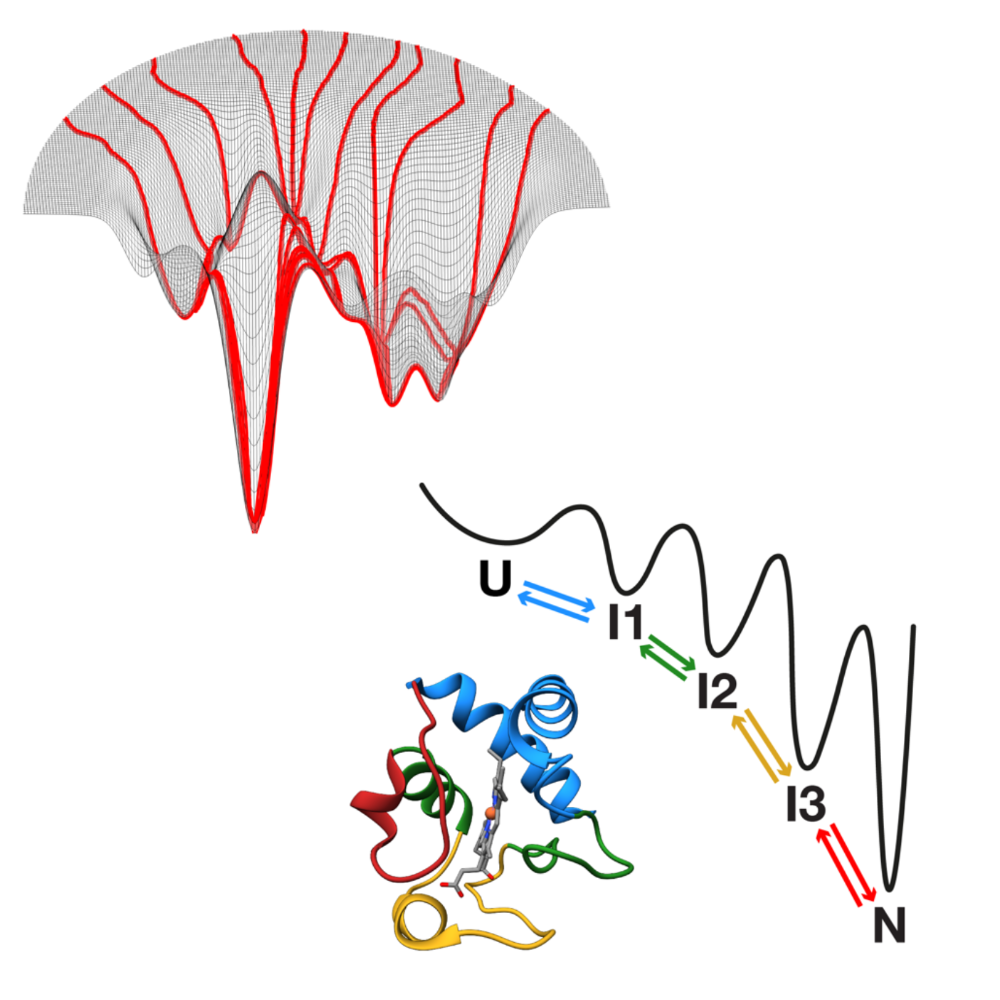
How proteins fold remains a central unsolved problem in biology. While the idea of a folding code embedded in the amino acid sequence was introduced more than 6 decades ago, this code remains undefined. While we now have powerful predictive tools to predict the final native structure of proteins, we still lack a predictive framework for how sequences dictate folding pathways. Two main conceptual models dominate as explanations of folding mechanism: the funnel model, in which folding proceeds through many alternative routes on a rugged, hyperdimensional energy landscape; and the foldon model, which proposes a hierarchical sequence of discrete intermediates. Recent advances on two fronts are now enabling folding studies in unprecedented ways. Powerful experimental approaches; in particular, single-molecule force spectroscopy and hydrogen (deuterium exchange assays) allow time-resolved tracking of the folding process at high resolution. At the same time, computational breakthroughs culminating in algorithms such as AlphaFold have revolutionized static structure prediction, opening opportunities to extend machine learning toward dynamics. Together, these developments mark a turning point: for the first time, we are positioned to resolve how proteins fold, why they misfold, and how this knowledge can be harnessed for biology and medicine.