Jesus Pineda defended his PhD thesis on November 11th, 2025. Congrats!
The defense took place in SB-H7 lecture hall, Institutionen för fysik, Johanneberg Campus, Göteborg, at 9:00.
Title: Inductive Biases for Efficient Deep Learning in Microscopy
Abstract: Deep learning has become an indispensable tool for the analysis of microscopy data, yet its integration into routine research remains uneven. Several factors contribute to this gap, including the limited availability of well-annotated datasets and the high computational demands of modern architectures. Microscopy introduces further challenges, as it spans diverse modalities and scales, from proteins to tissues, producing heterogeneous data that defy standardization. Generating reliable annotations also requires expertise and time, while unequal access to high-performance computing further widens the divide between well-resourced institutions and smaller laboratories.
This dissertation argues that the prevailing paradigm of scaling models with ever-larger datasets and computational resources yields diminishing returns for microscopy. Instead, it explores the role of inductive biases as a foundation for building models that are more data-efficient, computationally accessible, and scientifically meaningful. Inductive biases are structural assumptions embedded in model design that guide learning toward patterns aligned with the underlying problem. The first part of this work examines their central role in the advancement of modern deep learning and the diverse ways they shape model behavior.
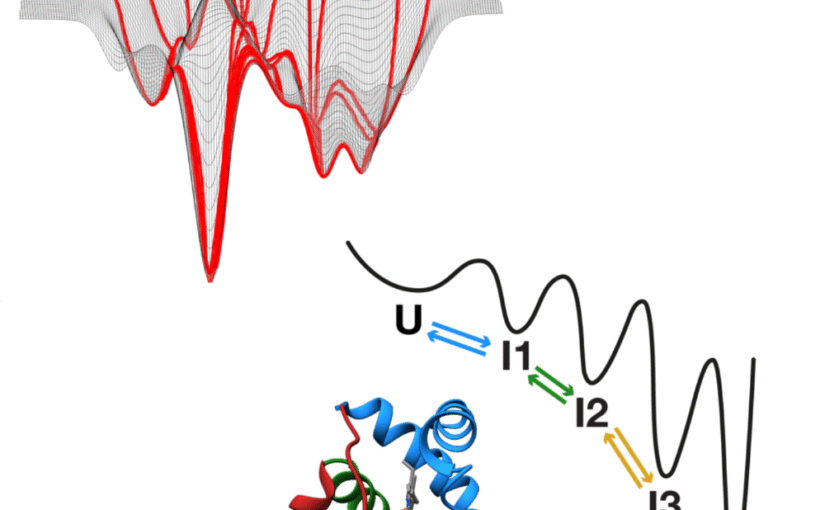
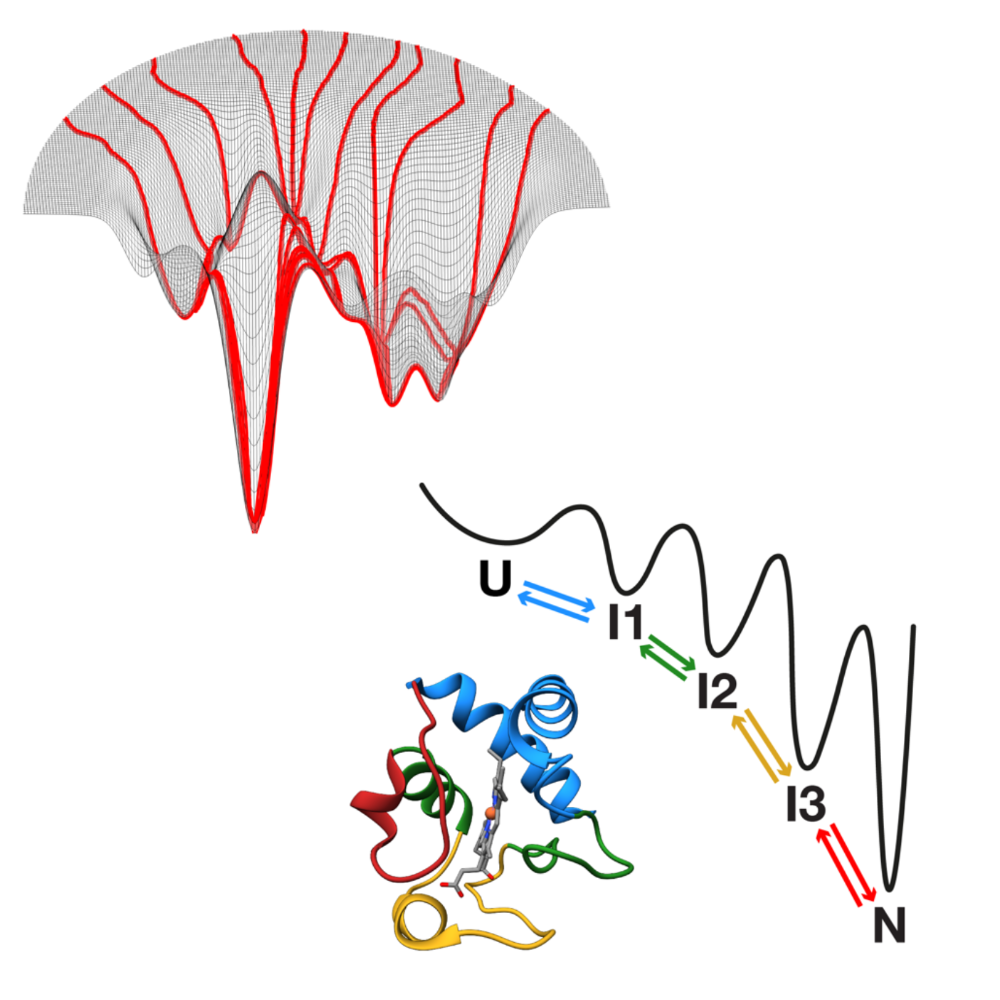
This potential is demonstrated through three case studies. First, MAGIK employs graph neural networks to analyze biological dynamics in time-lapse microscopy, uncovering local and global properties with high precision, even when trained on limited data. Next, MIRO leverages recurrent graph neural networks to process single-molecule localization datasets, improving the efficiency and reliability of clustering for variable biological structures and scales while retaining strong generalization with minimal supervision. Finally, GAUDI introduces a representation learning framework for characterizing biological systems, providing a physically meaningful representation space for interpretable and transferable analysis.
The findings presented here demonstrate that the integration of inductive biases provides a cohesive strategy to extend the reach of deep learning in the life sciences, enhancing accessibility and ensuring scientific utility under resource constraints.
Thesis: https://gupea.ub.gu.se/items/672c7946-51d6-4773-ad8c-35a3eed41499
Supervisor: Giovanni Volpe
Co-Supervisor: Carlo Manzo
Examiner: Raimund Feifel
Opponent: Anna Kreshuk
Committee: Juliette Griffié, Daniel sage, Daniel Persson
Alternate board member: Jonas Enger