Giovanni Volpe
DINAMO 2026
Franschhoek, South Africa
7 April 2026
Category: Presentation
Presentation by S. K. Mondal, online, 22 April, 2026
Optical Fiber Micro/Nano Axicon Tip: An Optical Imaging Platform
Samir K. Mondal
CSIR-CSIO, Chandigarh, India
Date: 22 April 2026
Time: 12:30
Place: Online on zoom
Optical fiber tip under structural modifications enhances light-matter interaction by focusing, collecting or modulating light in microscopic scale and combined with waveguide property, it emerges as a potential optical tool, especially for spectroscopic, endoscopic and imaging application. A chemical etching technique has been introduced to permanently modify the tip as Micro/Nano axicon, capable in generating structured beams. The optics of the axicons have been studied in detail and further used in optical imaging experiments, namely phase microscopy, photonic nanojet and nanoscopy. The seminar will highlight first-hand information about the probe and experiments addressing the above-mentioned application.
Short Bio
Dr. Mondal is Chief Scientist at CSIR-CSIO, Chandigarh. He earned his Ph.D. in Electronic Science and M.Sc. in Physics from the University of Calcutta. After postdoctoral research at the University of California, Irvine and the University of Minnesota, he joined Tyndall Research Institute, Ireland.
With over 25 years in optics and photonics, his work spans optical interconnects, photonic crystals, lasers, and fiber instrumentation. He leads research in optical fiber antennas, near-field optics, imaging, and plasmonics, aiming for sustainable photonics platforms.
He collaborates internationally and is known for pioneering micro/nano axicons on fiber tips. He has over 50 publications and serves as an editor and reviewer.
Poster by P. Dutta at the Protein Folding in Real Time Conference, Stockholm, 11th March 2026
Prakhar Dutta, Jiacheng Huang, Nazli Demirpehlivan, Thomas Catley, Sylvia Whittle, Carlo Manzo, Rahul Nagshi, Rachel Owen, Giovanni Volpe
Date: 11th March 2026
Time: 18:00 – 20:00
Place: Aula Medica, Karolinska Institute, Solna
Conference Protein Folding in Real Time, 11-13 March 2026, Stockholm, Sweden
Abstract: Atomic force microscopy (AFM) resolves biological structure and mechanics at high resolution, but produces vast, heterogeneous datasets that are often noisy and very time-consuming to analyse. Although deep learning could automate quality control, segmentation and feature extraction, adoption is limited by scarce ground-truth training data and high technical barriers for experimentalists. Here we present ASAP, an open-source tutorial and pipeline implemented in DeepTrack to provide a reproducible foundation for AI-enabled AFM. At the protein folding conference, a dual-pathway simulation for DNA, offering both molecular dynamics and rapid, non-MD geometries to generate perfect ground truth for segmentation training was presented. By consolidating simulation and learning into a single modular ecosystem, this work enables users to build upon our pipeline to optimize AFM workflows for more efficient data acquisition and robust processing.
Poster by Anqi Lyu at the Protein Folding in Real Time conference, Stockholm, 11 March 2026
Anqi Lyu, Maria Jesus Iglesias, Jochen Schwenk, Mathias Uhlén, Jacob Odeberg, Caroline Adiels, Lynn Butler
Date: 11 March 2026
Time: 18:00-20:00
Place: Aula Medica, Karolinska Institute, Solna
Conference Protein Folding in Real Time, 11-13 March 2026, Stockholm, Sweden
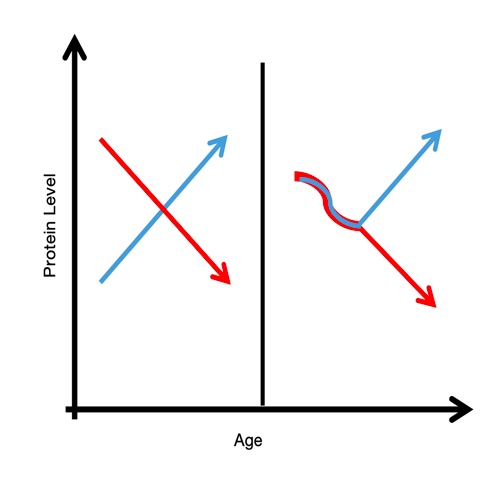
Aging is biologically heterogeneous, and chronological age alone cannot explain molecular variability across individuals. Previous studies have shown that disease states can reshape age-associated proteomic trajectories, leading to divergent molecular patterns over time. Here, we explore these dynamics by analyzing age-dependent changes in plasma protein abundance, focusing on differences between case and control conditions during aging. We identify opposing trends and nonlinear transitions, particularly in later life, highlighting critical periods of accelerated molecular change. Beyond descriptive patterns, our analysis emphasizes how disease modifies the underlying structure of aging trajectories, providing insights into the mechanisms of age-related divergence.
Poster by M. Granfors at the Protein Folding in Real Time conference, Stockholm, 11 March 2026
Mirja Granfors, Jakub Masaryk, Carlo Manzo, Markus Tamas, Giovanni Volpe
Date: 11th March 2026
Time: 18:00 – 20:00
Place: Aula Medica, Karolinska Institute, Solna
Conference Protein Folding in Real Time, 11-13 March 2026, Stockholm, Sweden
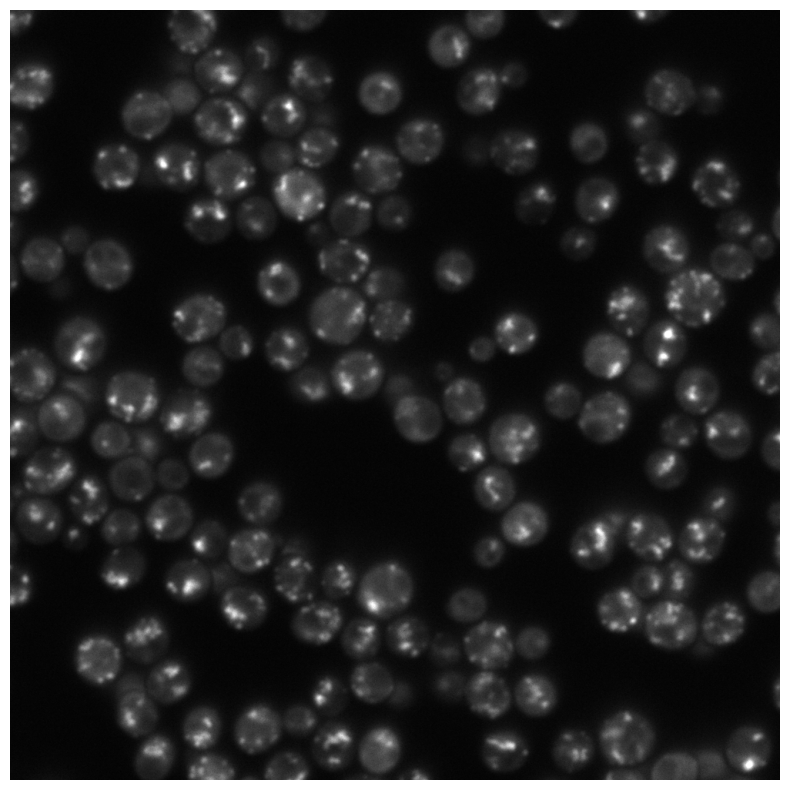
Arsenic is a toxic metal linked to serious diseases like cancer and neurodegeneration. One proposed mechanism of toxicity is that arsenic causes proteins to misfold and aggregate inside cells, but the dynamics and regulation of this process remain poorly understood. Using fluorescence microscopy data from living yeast cells, we are developing a machine learning approach to automatically detect, track, and analyze protein aggregate movement over time.
Poster by X. Zhang at at the Protein Folding in Real Time conference, Stockholm, 11 March 2026
Xinwen Zhang, Nora Haanaes, Berenice García Rodríguez, Giovanni Volpe, Janet Kumita and Daniel Midtvedt
Date: 11 March 2026
Time: 18:00-20:00
Place: Aula Medica, Karolinska Institute, Solna
Conference Protein Folding in Real Time, 11-13 March 2026, Stockholm, Sweden
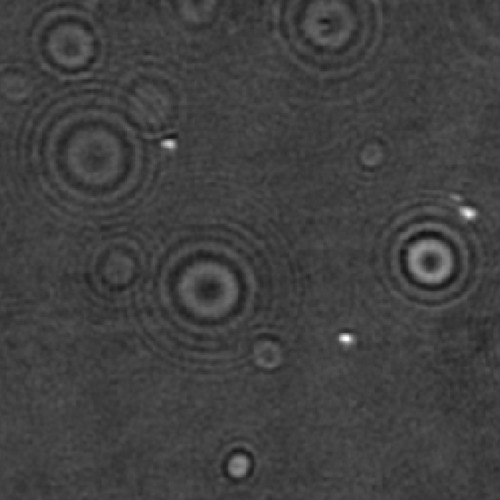
Biomolecular condensates formed through liquid–liquid phase separation (LLPS) play important roles in cellular organization, yet quantitative and label-free characterization of their physical properties remains challenging. In this work, we apply off-axis holographic microscopy to study a synthetic biomolecular condensate platform based on the LCD2-CTPR protein system. These proteins, composed of modular consensus-designed tetratricopeptide repeat (CTPR) domains fused to intrinsically disordered regions, undergo phase separation under varying salt concentrations. By incorporating short binding motifs such as ATG13 or Func1, the condensates can specifically recruit the autophagy-related protein LC3. Using label-free quantitative phase measurements, we analyze changes in condensate optical radius and refractive index during LC3 recruitment and over time. Our results show measurable variations in condensate size and optical properties, highlighting the sensitivity of these systems to compositional changes. This work demonstrates the applicability of holographic microscopy for quantitative characterization of synthetic biomolecular condensates and provides a framework for studying protein phase separation in a non-invasive manner.
Poster by E. A. Duta Costache at the Protein Folding in Real Time conference, Stockholm, 11 March 2026
Eduard Duta Costache, Benjamin Girault
Date: 11 March 2026
Time: 18:00-20:00
Place: Aula Medica, Karolinska Institute, Solna
Conference Protein Folding in Real Time, 11-13 March 2026, Stockholm, Sweden
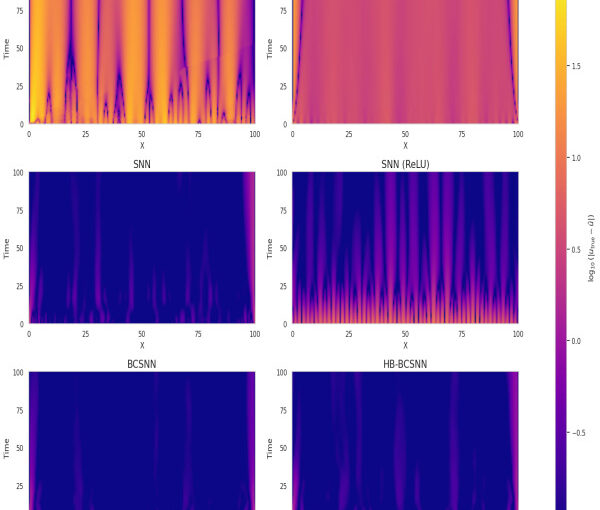
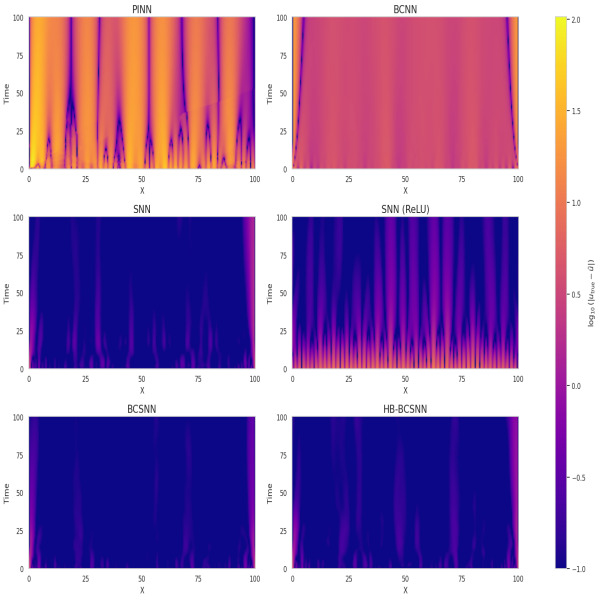
Physics-Informed Neural Networks (PINNs) have emerged as a promising method for solving Partial Differential Equations (PDEs) by combining data-driven learning with physical laws. However, the spectral bias and optimization challenges limit their efficacy. This work investigates these issues and whether the advantages of classical spectral methods translate to the non-convex neural network optimization landscape. We show that gradient imbalance greatly affect learning and we study the Hessian conditioning under different settings. Our results indicate that spectral priors stabilize training, reduce error, and improve parameter efficiency. We also identify that learnable-basis models act as implicit regularizers under sparse sampling.
Poster by Sreekanth K Manikandan at the Protein Folding in Real Time conference, Stockholm, 11 March 2026
Sreekanth K Manikandan
Date: 11 March 2026
Time: 18.00-20.00
Place: Aula Medica, Stockholm Sweden
Conference Protein Folding in Real Time, 11-13 March 2026, Stockholm, Sweden
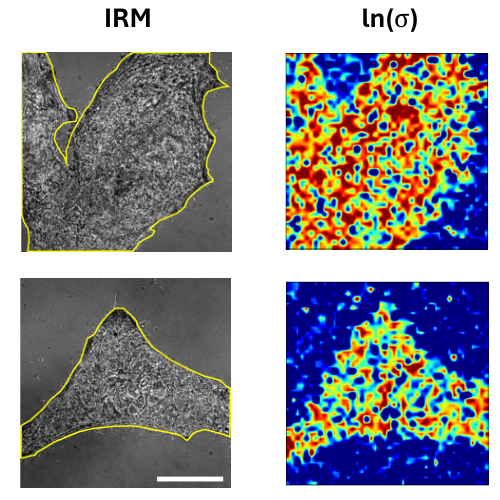
Identifying whether a process is in equilibrium, quantifying its distance from equilibrium, and constructing optimal reduced descriptions of non-equilibrium dynamics remain central challenges in the study of living matter. Here, we discuss how data-driven approaches grounded in stochastic thermodynamics enable these features to be inferred directly from experimental data. In particular, we show how entropy production can be localized in space and time, and how maximally dissipative coordinates emerge as effective low-dimensional descriptions of non-equilibrium processes. We highlight applications to experimental biophysical systems and discuss key challenges and limitations.
Poster by M. Selin at the Protein Folding in Real Time conference, Stockholm, 11 March 2026

Martin Selin, Antonio Ciarlo, Giuseppe Pesce, Lars Bengtsson, Joan Camunas-Soler, Vinoth Sundar Rajan, Fredrik Westerlund, L. Marcus Wilhelmsson, Isabel Pastor, Felix Ritort, Steven B. Smith, Carlos Bustamante, and Giovanni Volpe
Date: 11th March 2026
Time: 18:00 – 20:00
Place: Aula Medica, Karolinska Institute, Solna
Conference Protein Folding in Real Time, 11-13 March 2026, Stockholm, Sweden
Single-molecule studies are vital for understanding fundamental biological processes, including protein folding, DNA transcription, and replication. However, performing these experiments manually on individual molecules is notoriously time-consuming and costly. To address this challenge, we have developed a fully autonomous single-molecule force spectroscopy platform by integrating a custom-built optical tweezers instrument with real-time deep-learning-based image analysis and adaptive control protocols. Our system achieves human-level throughput in terms of experiments per hour while remaining robust enough to operate continuously for hours without intervention. We demonstrate the versatility of our platform by having it perform DNA pulling experiments fully autonomously. By making the software open source we democratize high-throughput data collection in single-molecule biophysics, paving the way for merging single-molecule studies with large-scale, data-driven approaches—ultimately enabling new insights into the dynamic, transient states of complex biological systems.
Poster by A. Schiano di Colella at the Protein Folding in Real Time conference, Stockholm, 11 March 2026
Andrea Schiano di Colella, Antonio Ciarlo, Mats Granath, Giovanni Volpe
Date: 11 March 2026
Time: 18:00 – 20:00
Place: Aula Medica, Karolinska Institutet, Stockholm, Sweden
Conference: Protein Folding in Real Time
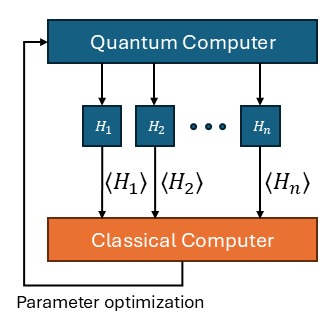
Quantum computing is a field of study that aims to exploit quantum mechanical effects for the purposes of computation. Due to the intrinsic capacity of qubits of efficiently represent an exponentially large configuration space, quantum computation has been identified as a promising candidate for complex physical chemistry simulations, including investigating the dynamics of protein folding. This work illustrates the use of quantum computing for variational problems, and the use of alternative training methods such as genetic algorithms to avoid the “barren plateau” phenomenon, which prevents the training of general quantum circuits by means of the usual gradient descent.