Zhilei Xu, Mite Mijalkov, Jiawei Sun, Yu-Wei Chang, Arianna Sala, Giovanni Volpe, Mario Severino, Mattia Veronese, Sara Garcia-Ptacek, Joana B. Pereira, for the Alzheimer’s Disease Neuroimaging Initiative
Alzheimer’s & Dementia 22, e71310 (2026)
DOI: 10.1002/alz.71310
INTRODUCTION
Mapping individual differences is crucial to improve personalized medicine approaches in Alzheimer’s disease (AD), which is characterized by strong inter-individual variability in the accumulation patterns of tau and amyloid beta pathology.
METHODS
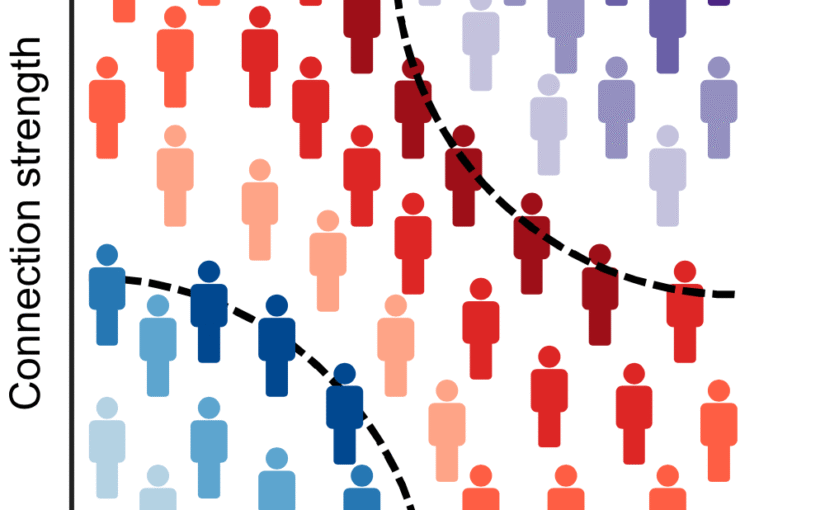
We assess the progression of AD across the disease continuum by building individual molecular connectomes using longitudinal positron emission tomography (PET) data.
RESULTS
We demonstrate that these connectomes constitute a unique fingerprint, capable of identifying a single individual from a large group of subjects. Alterations in the connectomes discriminate different diagnostic groups and predict cognitive decline to a higher extent than conventional PET measures. We introduce a novel gene-specific transcription network analysis that linked individual tau and amyloid connectomes to a common transcriptomic profile of apoptosis, with the tau connectome being specifically related to pyrimidine metabolism, and the amyloid connectome to histone acetylation.
DISCUSSION
Individual molecular connectome mapping provides a novel and sensitive framework to monitor AD progression.
Highlights
- Individual molecular connectomes constitute a unique fingerprint, capable of identifying a single individual from a large group of subjects.
- Alterations in individual molecular connectomes significantly increase both across the Alzheimer’s disease (AD) continuum and over time.
- Alterations in individual molecular connectomes discriminate different diagnostic groups and predict cognitive decline to a higher extent than conventional positron emission tomography measures.
- Susceptibilities of individual tau and amyloid connectomes to AD are linked to a common transcriptomic profile of apoptosis, with the tau connectome being specifically related to pyrimidine metabolism, and the amyloid connectome to histone acetylation.