Giovanni Volpe
Seminar
Vi2 Seminar (Visual Information and Interaction)
University of Uppsala
17 May 2021, 14:15 CEST
Online
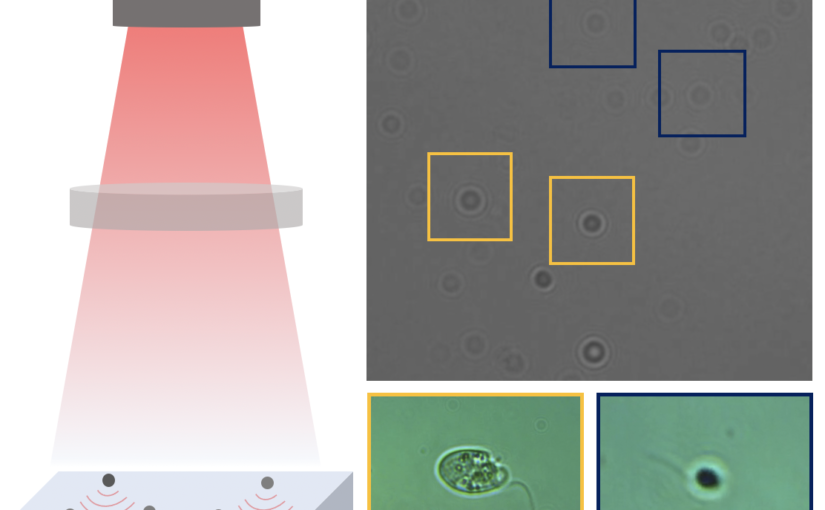
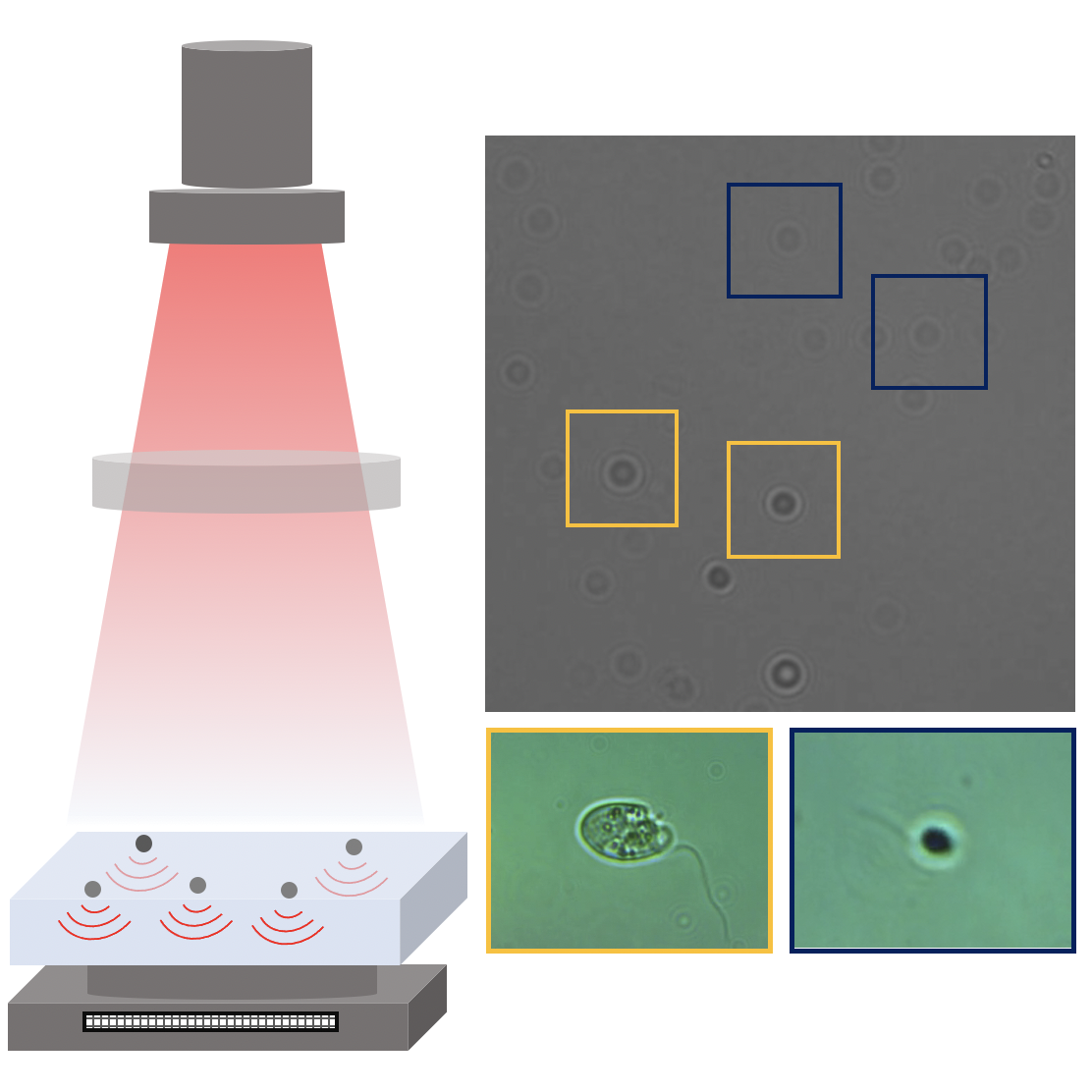
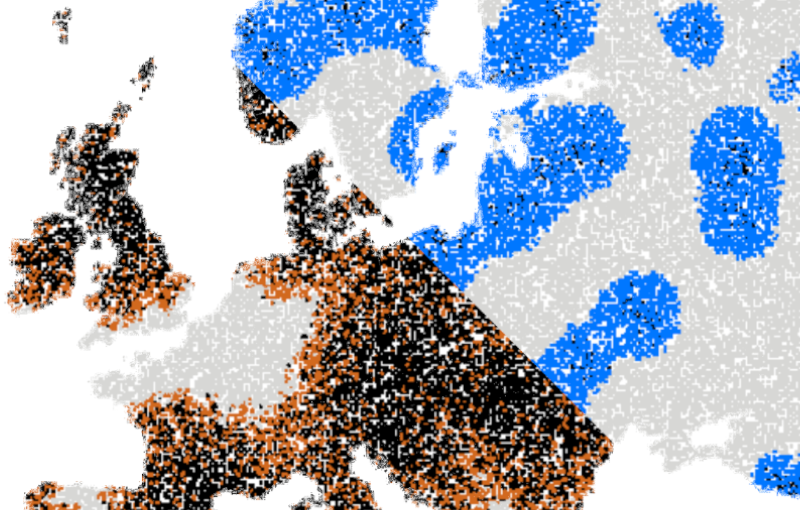
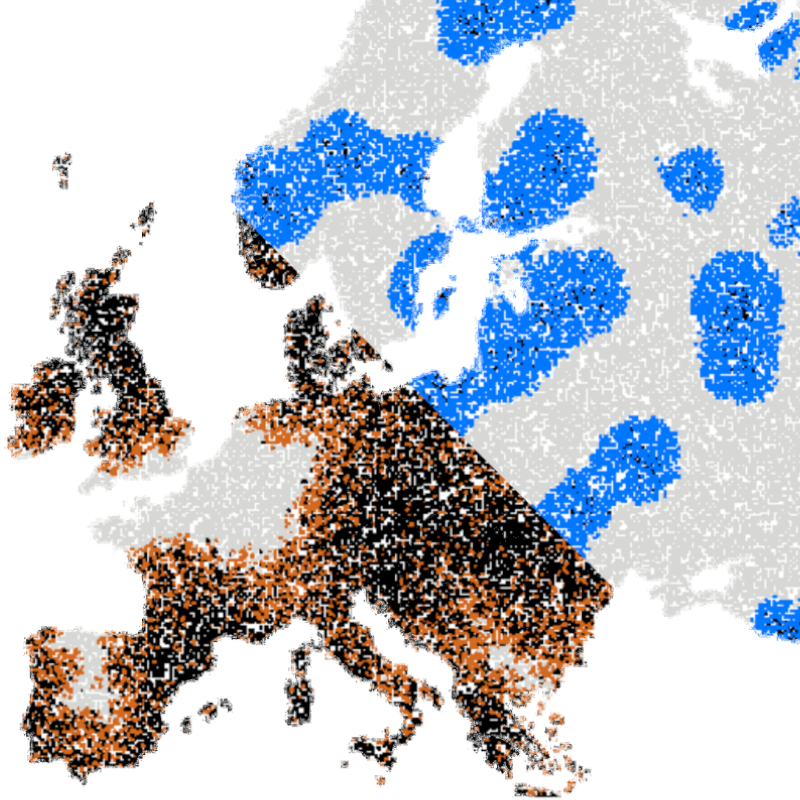
Video microscopy has a long history of providing insights and breakthroughs for a broad range of disciplines, from physics to biology. Image analysis to extract quantitative information from video microscopy data has traditionally relied on algorithmic approaches, which are often difficult to implement, time consuming, and computationally expensive. Recently, alternative data-driven approaches using deep learning have greatly improved quantitative digital microscopy, potentially offering automatized, accurate, and fast image analysis. However, the combination of deep learning and video microscopy remains underutilized primarily due to the steep learning curve involved in developing custom deep-learning solutions. To overcome this issue, we introduce a software, DeepTrack 2.0, to design, train and validate deep-learning solutions for digital microscopy. We use it to exemplify how deep learning can be employed for a broad range of applications, from particle localization, tracking and characterization to cell counting and classification. Thanks to its user-friendly graphical interface, DeepTrack 2.0 can be easily customized for user-specific applications, and, thanks to its open-source object-oriented programming, it can be easily expanded to add features and functionalities, potentially introducing deep-learning-enhanced video microscopy to a far wider audience.
Bio: Giovanni Volpe is Professor at the Physics Department at the University of Gothenburg (Gothenburg, Sweden), where he has been leading the Soft Matter Lab since 2016. He has established a strong research group of 18 people (3 postdocs, 12 PhD students, 3 Master students, http://www.softmatterlab.org ) with an externally-funded, ambitious and interdisciplinary research program that combines soft condensed matter, optical manipulation, nanotechnology, and machine learning. He has attracted external funding exceeding 6M €, including several national and European grants such as the ERC-StG ComplexSwimmers (2016-2021) and the ERC-CoG MAPEI (2021-2026). He is a co-funder of the startup companies Lucero Bio and IFLAI.
Link: Vi2 Seminars (zoom link included in the webpage)