Martin Selin
Single-Molecule Sensors and NanoSystems International Conference – S3IC 2023
23 November 2023, 11:51 (CET)
Since their invention by Ashkin et al. in the 1980s, optical tweezers have evolved into an indispensable tool in physics, especially in biophysics, with applications spanning from cell sorting to stretching single DNA strands. By the 2000s, commercial systems became available. Nevertheless, owing to their unique requirements, many labs prefer to construct their own, often drawing inspiration from existing designs.
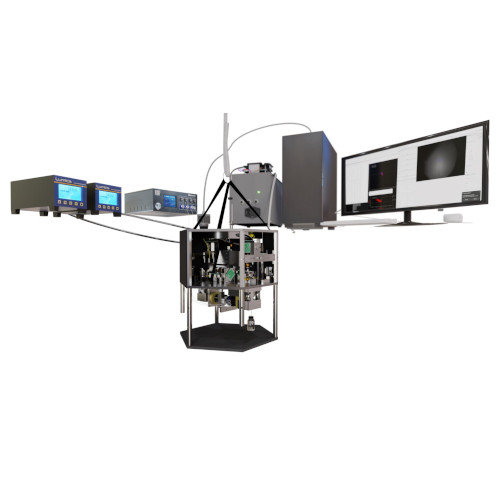
A prominent optical tweezers design is the “miniTweezers” system, pioneered by Bustamante’s group in the late 1990s. This system has been widely adopted globally for force spectroscopy experiments on single molecules, including DNA, proteins, and RNA.
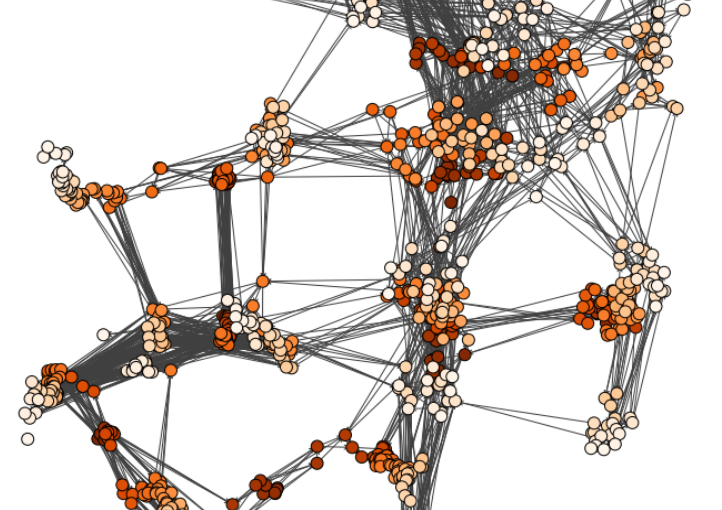
In this presentation, we unveil an advanced iteration of the miniTweezers. By enhancing its control and acquisition capabilities, we’ve augmented its versatility, enabling new experiment types. A significant breakthrough is the integration of real-time image feedback, which paves the way for automated procedures via deep learning-based image analysis, the first of which we demonstrate in this presentation.
We showcase this system’s capabilities through three distinct experiments:
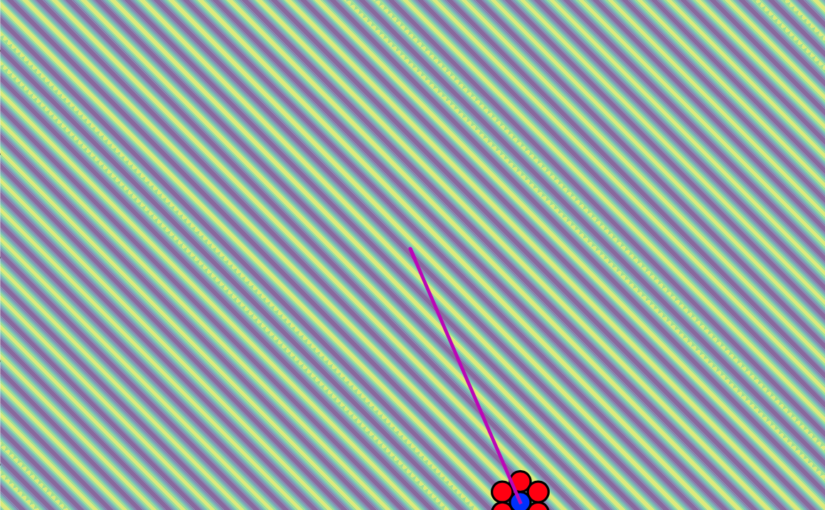
- A pulling experiment on a λ-DNA strand. By tethering DNA between two polystyrene beads – one anchored in a micropipette and the other manipulated by the tweezer – we illustrate near-complete automation, with the system autonomously handling bead trapping, attachment of the DNA and the pulling procedure.
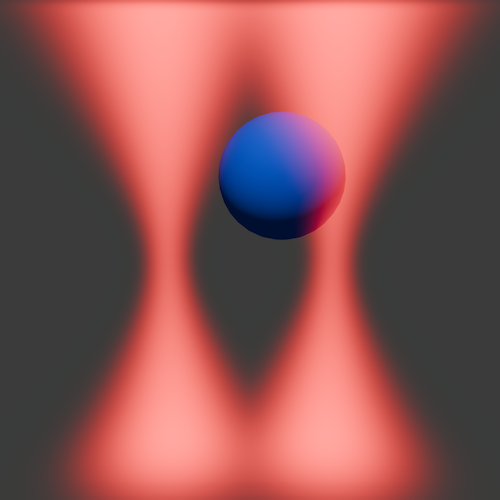
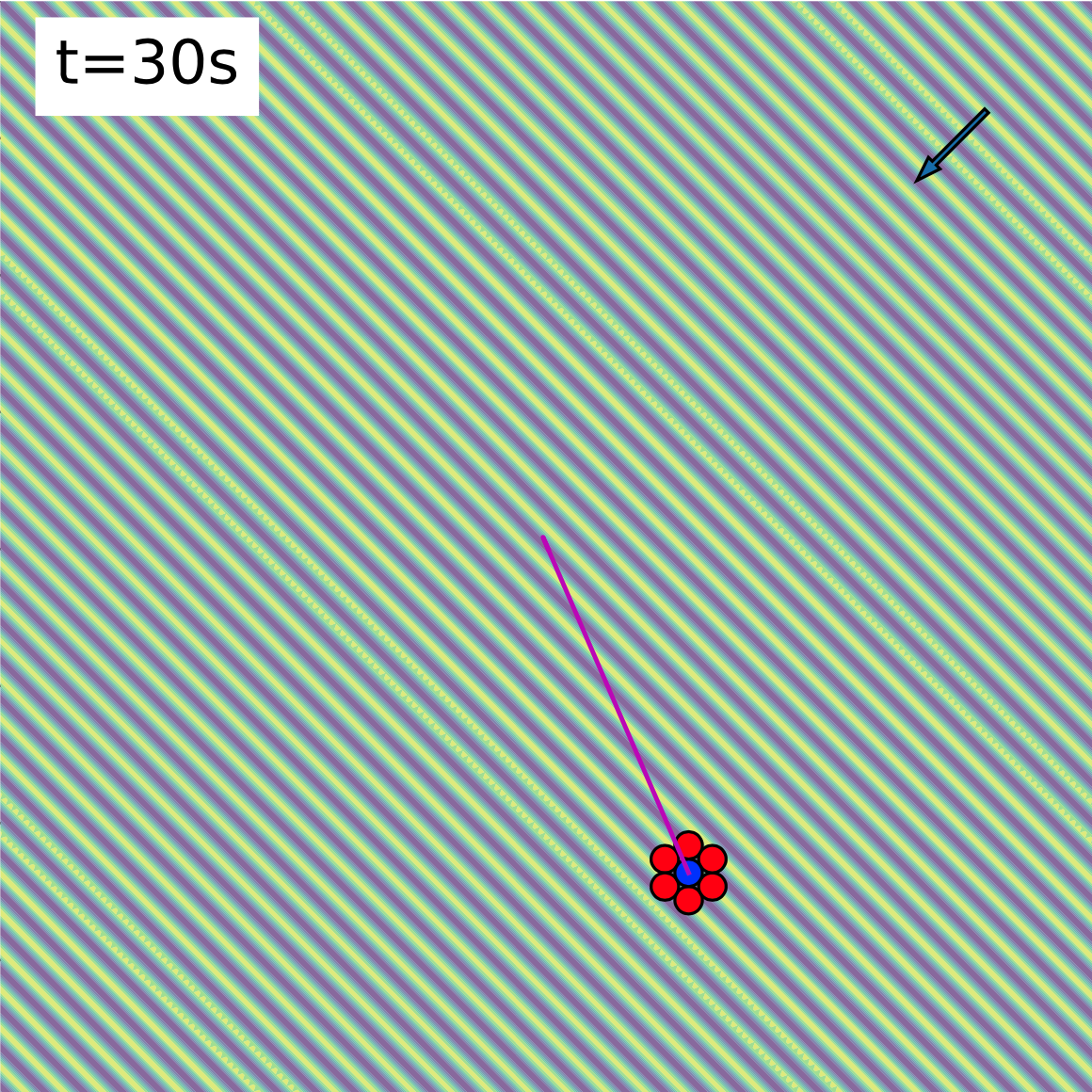
- An exploration of Coulomb interactions between charged particles. Here, one particle remains in a micropipette, while the other orbits the stationary bead, providing a 3D map of the interaction.
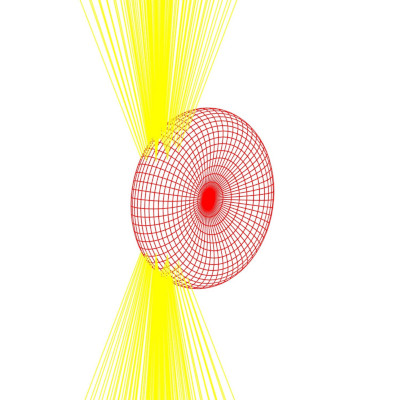
- A non-contact stretching experiment on red blood cells is conducted under low osmotic pressure conditions. Modulating the laser power induces cell elongation along the laser’s propagation direction. By correlating this elongation with the optical force exerted by the lasers, we present a simple and non-invasive method to measure membrane rigidity.
In summary, these advancements mark a significant leap in the capabilities and applications of optical tweezers in biophysics. As we push the boundaries of automation and precision, we envision a future where such instruments can unravel even more intricate molecular interactions and cellular mechanics, setting the stage for groundbreaking discoveries.