Henrik K. Moberg, Bohdan Yeroshenko, Joachim Fritzsche, David Albinsson, Barbora Spackova, Daniel Midtvedt, Giovanni Volpe, Christoph Langhammer
Nature Communications 17, 2533 (2026)
DOI: 10.1038/s41467-026-70514-z
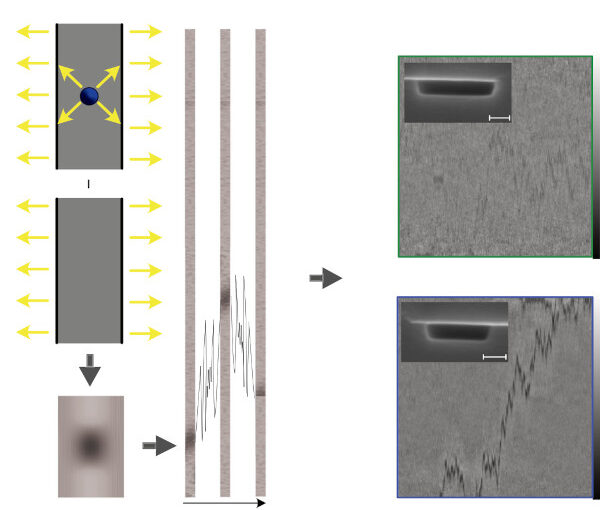
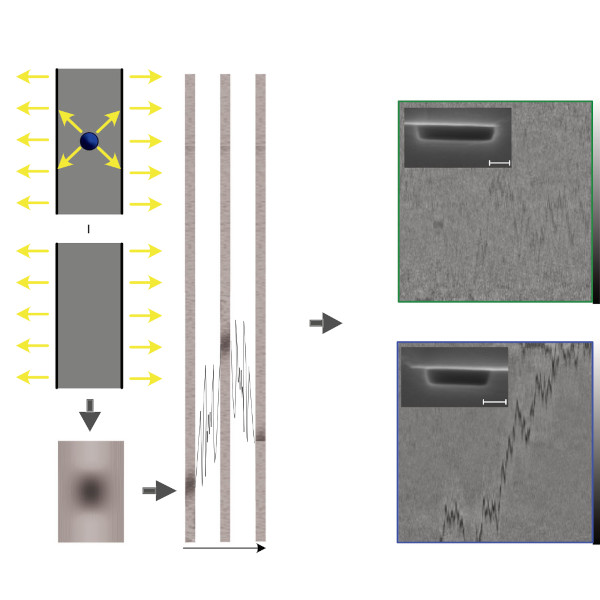
Nanofluidic scattering microscopy characterizes single molecules in subwavelength nanofluidic channels label-free, using the interference of visible light scattered by the molecule and nanochannel. It determines a molecule’s hydrodynamic radius by tracking its diffusion trajectory and its molecular weight by analyzing its scattering intensity along that trajectory. However, using standard analysis algorithms, it is limited to characterization of proteins larger than ≈ 60 kDa. Here, we push this limit by one order of magnitude to below ≈ 6 kDa molecular weight and ≈ 1.5 nm hydrodynamic radius — as we exemplify on the peptide hormone insulin — by using ultrasmall nanofluidic channels and by analyzing the data with a hierarchical vision transformer. When we benchmark this approach against the theoretical limit set by the Cramér–Rao Lower Bound, we find that it can be approached with sufficiently long molecular trajectories. This enables quantitative label-free single-molecule microscopy for biologically relevant families of sub-10-kDa molecules, such as cytokines, chemokines and peptide hormones.