DeepTrack: A comprehensive deep learning framework for digital microscopy
Benjamin Midtvedt, Saga Helgadottir, Aykut Argun, Daniel Midtvedt, Giovanni Volpe
Click here to see the slides.
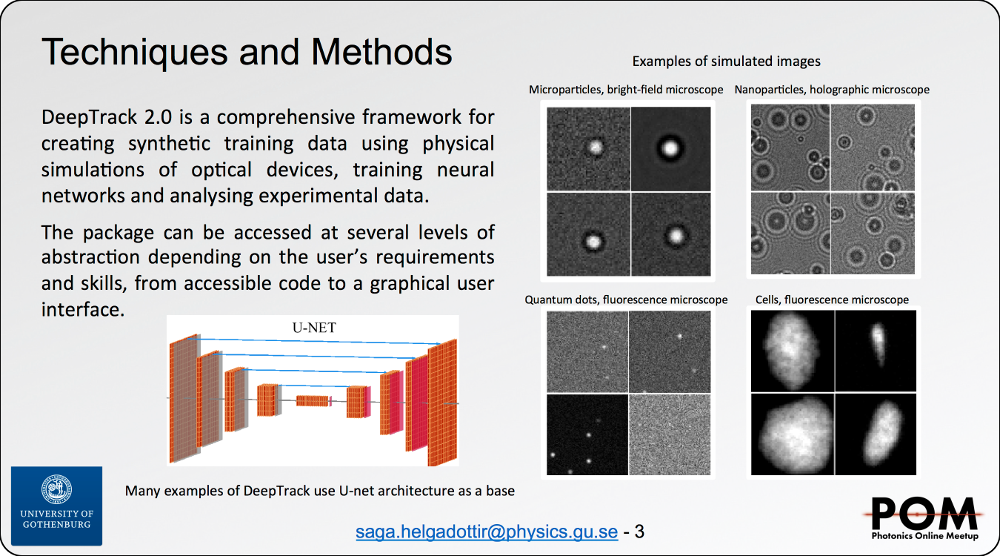
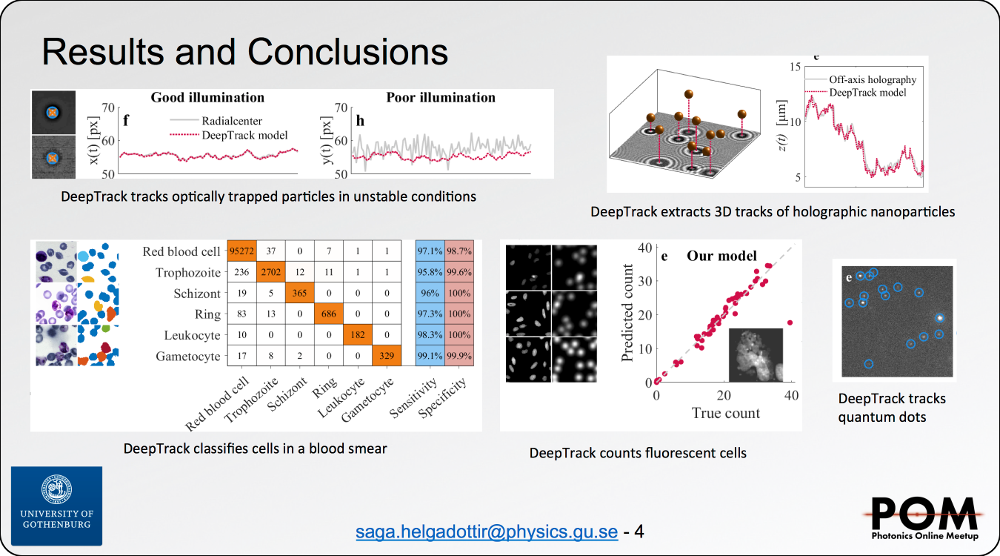
Despite the rapid advancement of deep learningmethods for image analysis, they remain under-utilized for the analysis of digital microscopy images. State of the artmethods require expertise in deep learning to implement, disconnecting the development of new methods from end-users. The packages that are available are typically highly specialized, diicult to reappropriate and almost impossible to interface with other methods. Finally, it is prohibitively difficult to procure representative datasets with corresponding labels. DeepTrack is a deep learning framework targeting optical microscopy, designed to account for each of these issues. Firstly, it is packaged with an easy-to-use graphical user interface, solving standard microscopy problems with no required programming experience. Secondly, it provides a comprehensive programming API for creating representative synthetic data, designed to exactly suit the problem. DeepTrack images samples of refractive index or flourophore distributions using physical simulations of customizable optical systems. To accurately represent the data to be analyzed, DeepTrack supports arbitrary optical aberration and experimental noise. Thirdly, many standard deep learning methods are packaged with DeepTrack, including architectures such as U-NET, and regularization techniques such as augmentations. Finally, the framework is fully modular and easily extendable to implement new methods, providing both longevity and a centralized foundation to deploy new deep learning solutions. To demonstrate the versatility of the framework,we show a few typical use-cases, including cell-counting in dense biological samples, extracting 3-dimensional tracks from 2-dimensional videos, and distinguishing and tracking microorganisms in bright-field videos.
Poster Session
Time: June 22nd 2020
Place: Twitter and virtual reality
POM Conference
Link: POM
Time: June 25th 2020
Place: Online