Jiacheng Huang, Nazli Demirpehlivan, Prakhar Dutta, Rahul Nagshi, Thomas Catley, Sylvia Whittle, Carlo Manzo, Rachel Owen, Giovanni Volpe
Date: 11th March 2026
Time: 18:00 – 20:00
Place: Aula Medica, Karolinska Institute, Solna
Conference Protein Folding in Real Time, 11-13 March 2026, Stockholm, Sweden
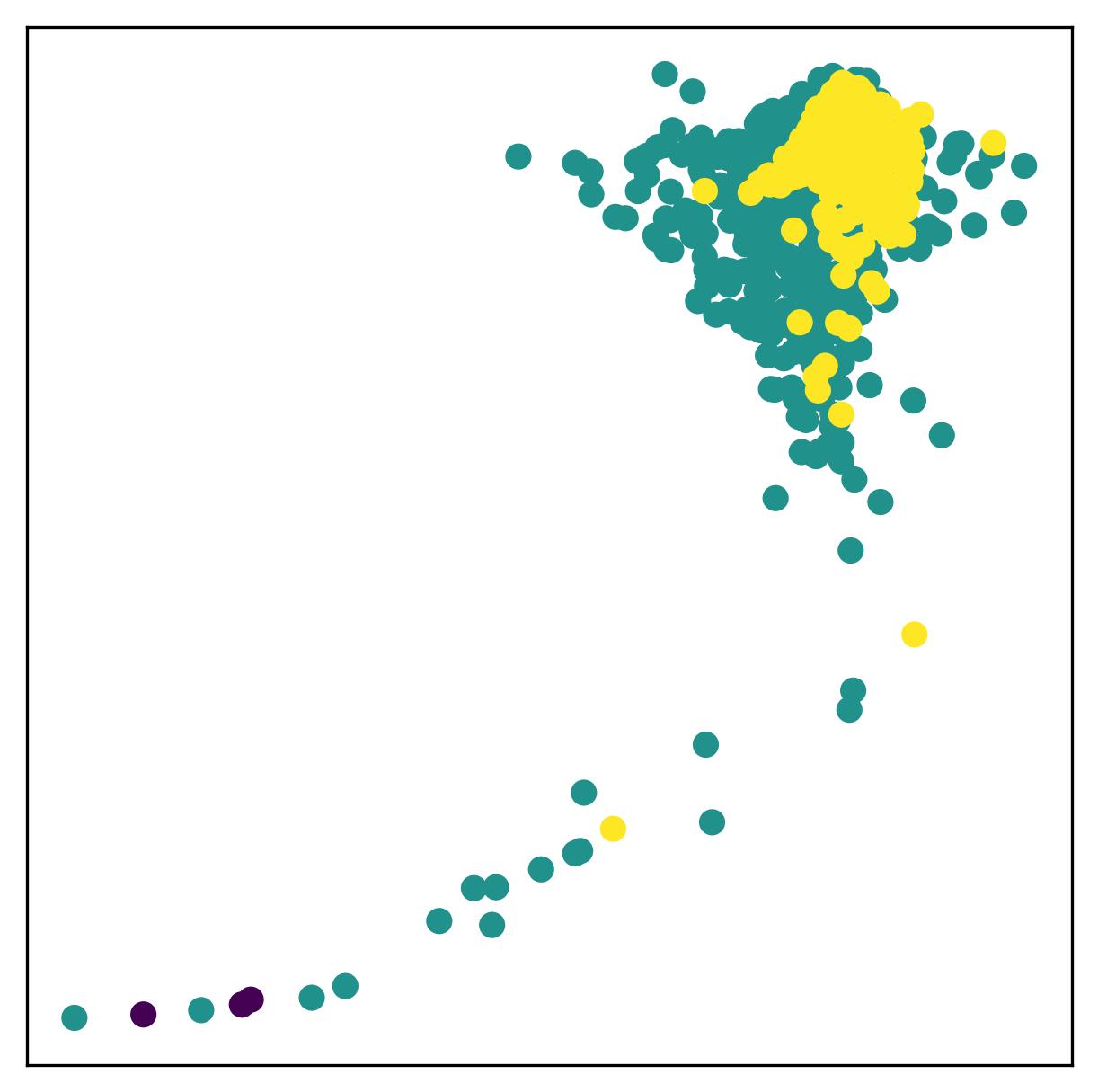
Atomic Force Microscopy (AFM) force spectroscopy is widely used to probe the mechanical properties and interactions of biological samples at the nanoscale, including living cells. However, large datasets generated during AFM measurements often contain curves affected by experimental artifacts such as poor tip–sample contact, noise, or instrumental instability. These low-quality force curves can significantly affect downstream analysis and typically require time-consuming manual inspection. In this work, we propose a machine learning–based data quality control framework for AFM force spectroscopy using a self-supervised approach.
