Mirja Granfors, Jakub Masaryk, Carlo Manzo, Markus Tamas, Giovanni Volpe
Date: 11th March 2026
Time: 18:00 – 20:00
Place: Aula Medica, Karolinska Institute, Solna
Conference Protein Folding in Real Time, 11-13 March 2026, Stockholm, Sweden
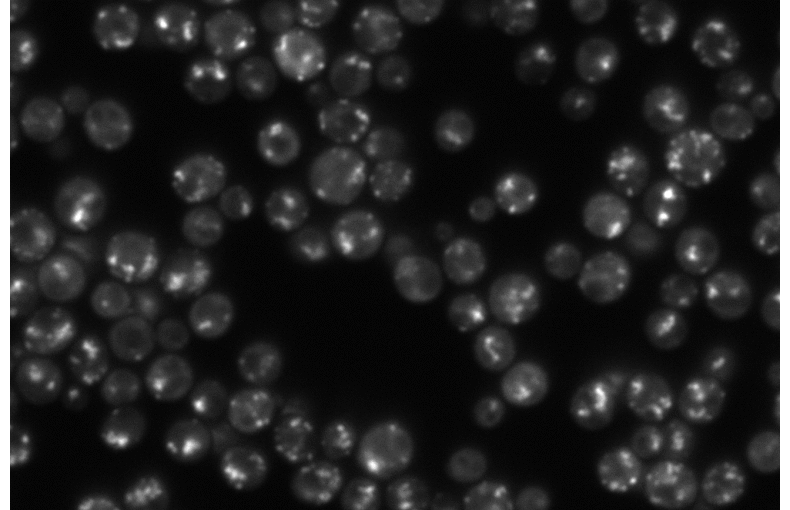
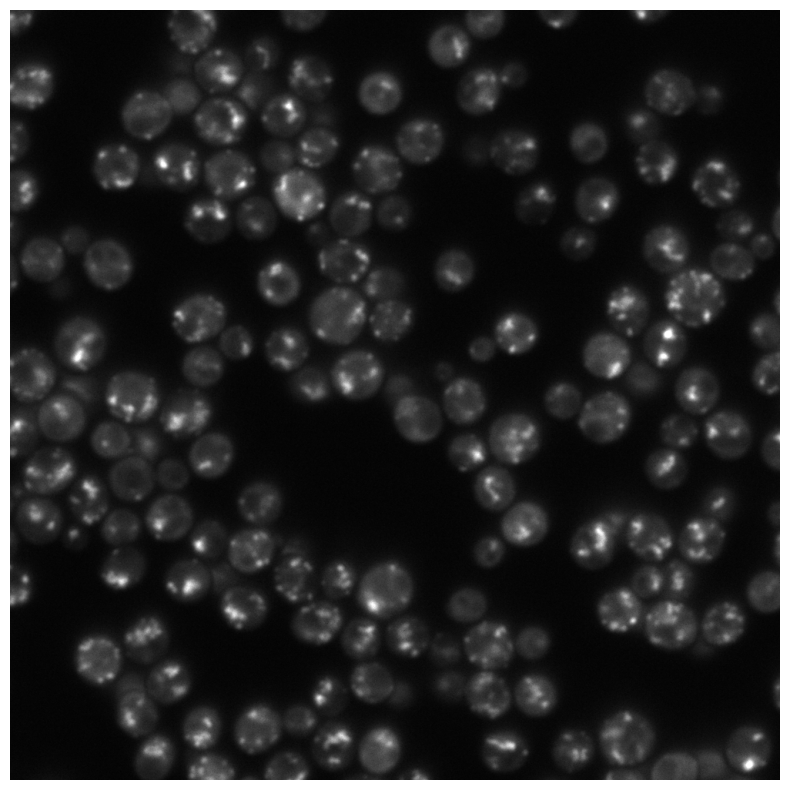
Arsenic is a toxic metal linked to serious diseases like cancer and neurodegeneration. One proposed mechanism of toxicity is that arsenic causes proteins to misfold and aggregate inside cells, but the dynamics and regulation of this process remain poorly understood. Using fluorescence microscopy data from living yeast cells, we are developing a machine learning approach to automatically detect, track, and analyze protein aggregate movement over time.