Laura Natali
Date: 22 August 2023
Time: 5:30 PM PDT
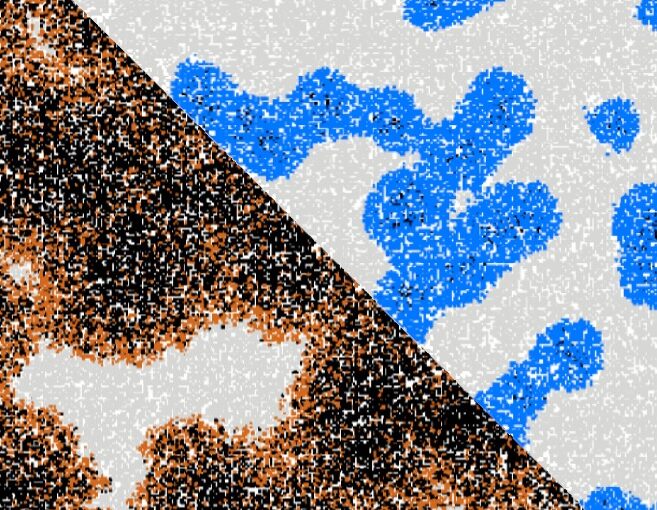
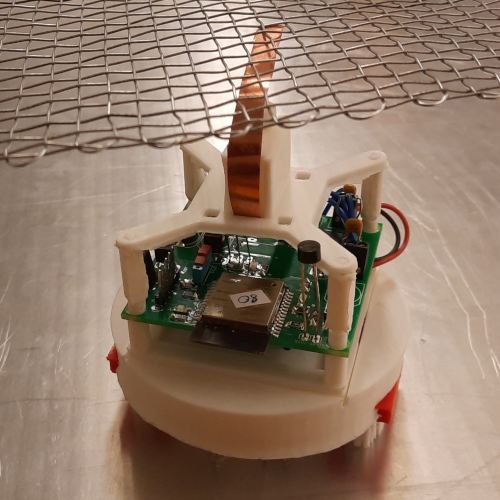
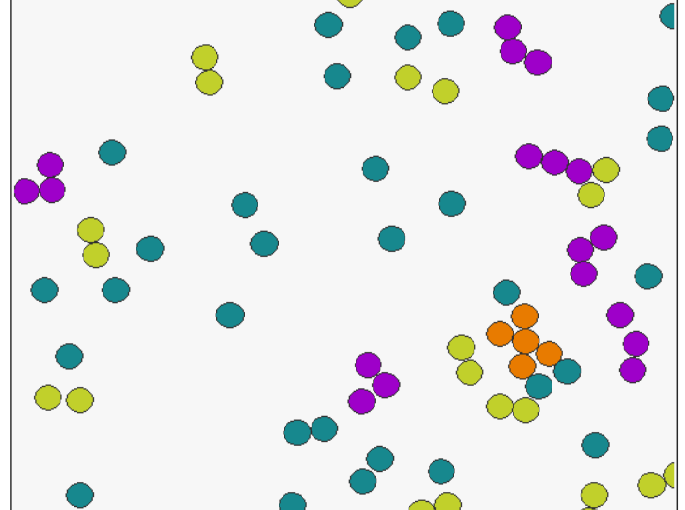
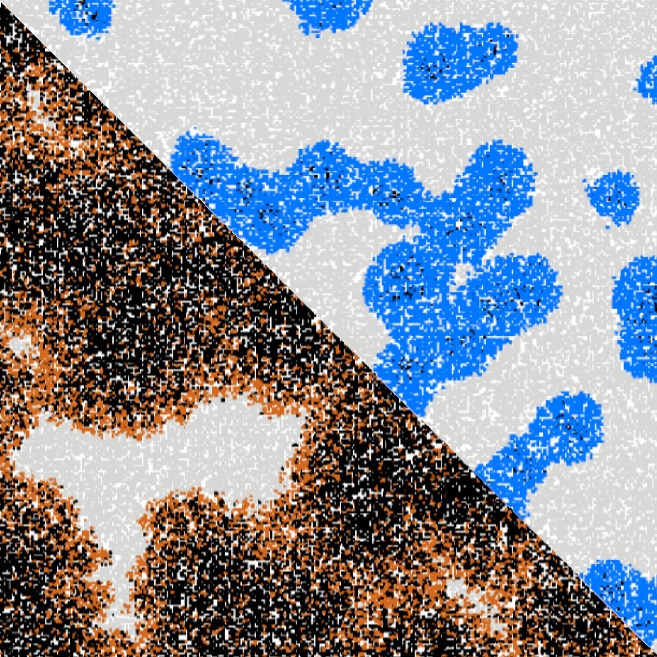
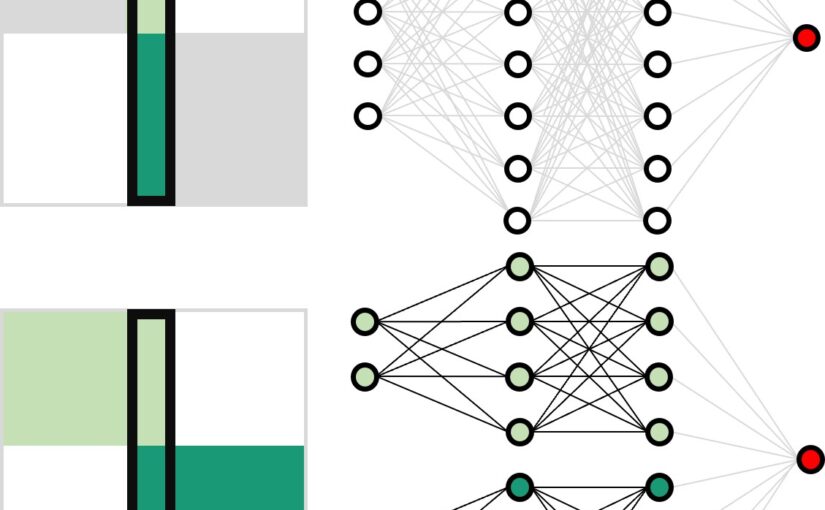
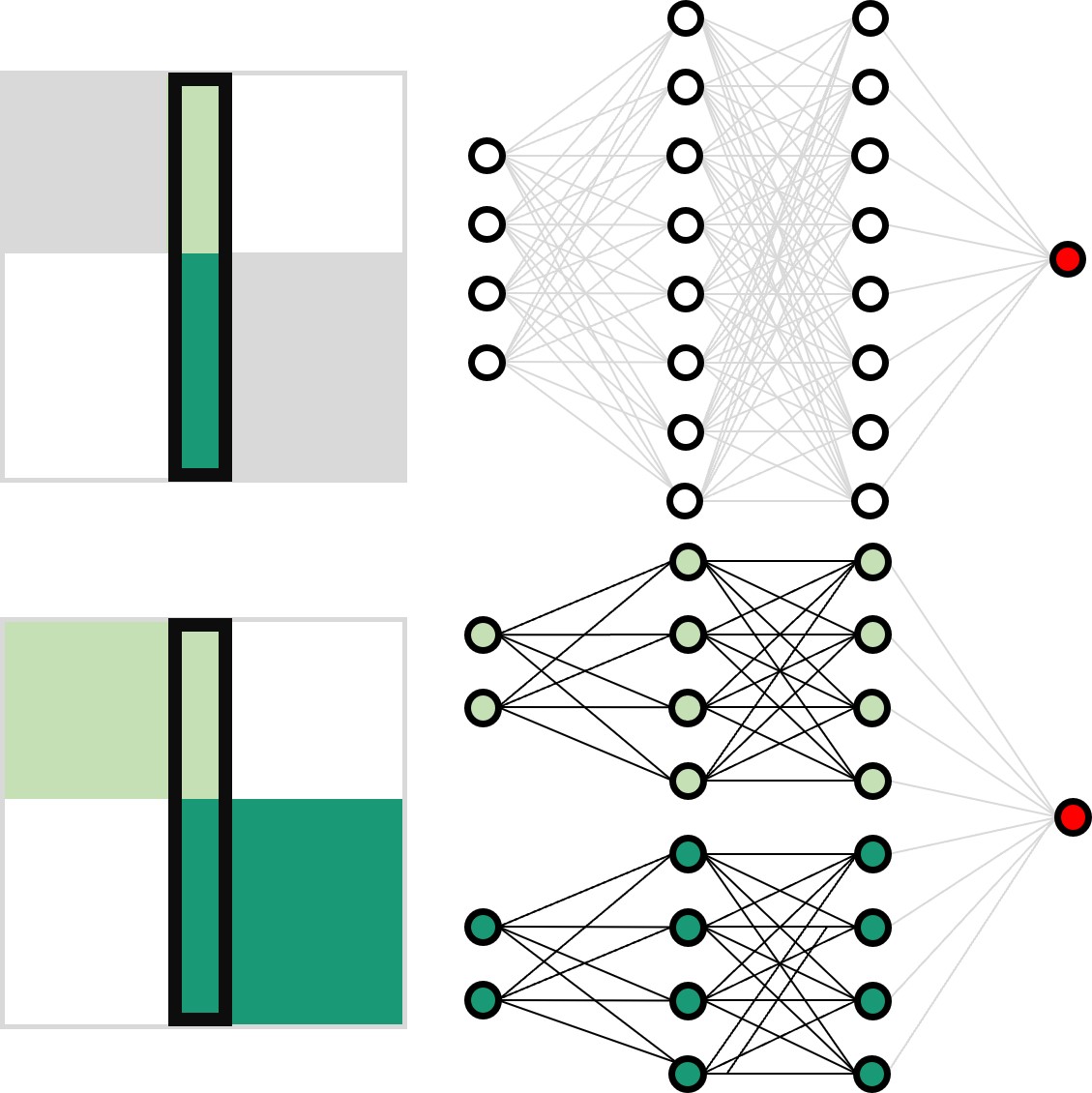
Artificial neural networks have limitations compared to biological counterparts as the latter can dynamically change connections and recover from damage. To simplify the study of connections evolving over time, we propose using programmable robots as a swarm to perform supervised learning and autonomously restructure the network. This experimental setup offers a way to study the evolution of connections in a simplified system while addressing the complexity of biological neurons. It has the potential to yield insights into the functioning of biological neural networks while providing a practical application in solving tasks.