Jesús Pineda, Benjamin Midtvedt, Harshith Bachimanchi, Sergio Noé, Daniel Midtvedt, Giovanni Volpe, and Carlo Manzo
Submitted to ISMC 2022
Date: 19 September 2022
Time: 13:40 (CEST)
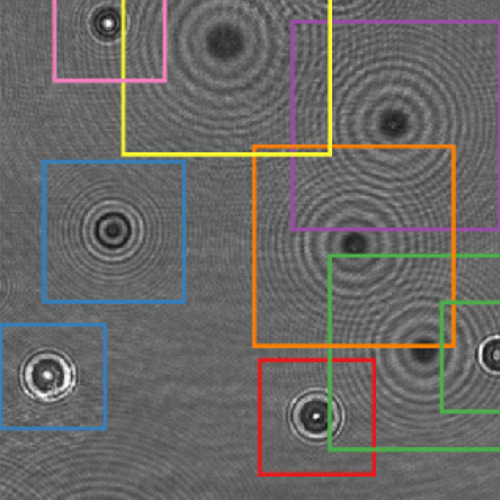
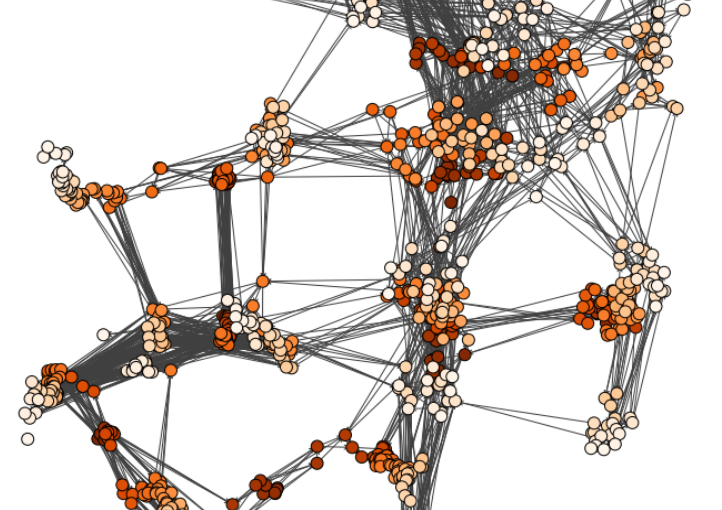
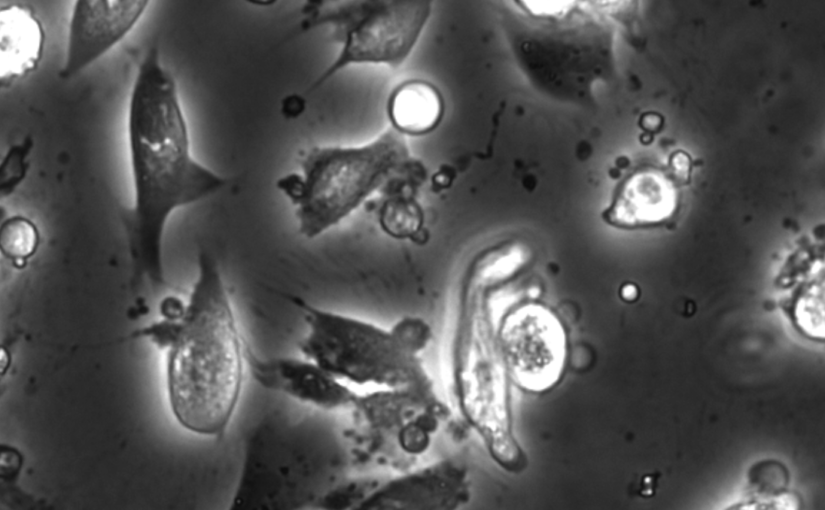
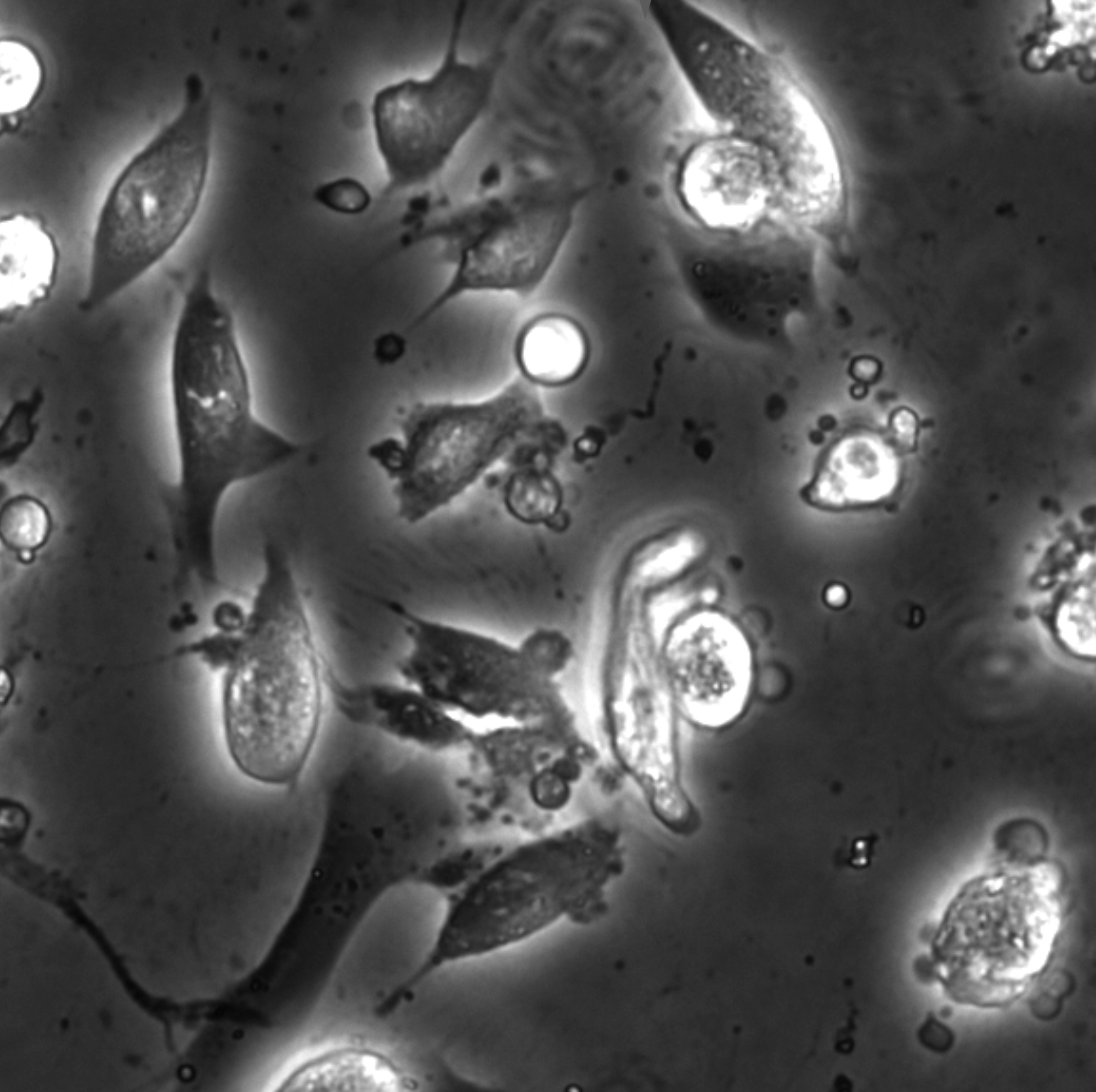
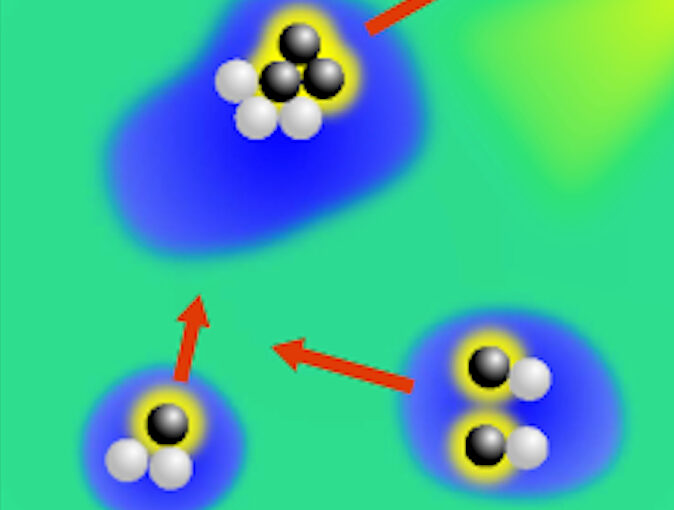
The characterization of dynamical processes in living systems provides important clues for their mechanistic interpretation and link to biological functions. Thanks to recent advances in microscopy techniques, it is now possible to routinely record the motion of cells, organelles, and individual molecules at multiple spatiotemporal scales in physiological conditions. However, the automated analysis of dynamics occurring in crowded and complex environments still lags behind the acquisition of microscopic image sequences. Here, we present a framework based on geometric deep learning that achieves the accurate estimation of dynamical properties in various biologically-relevant scenarios. This deep-learning approach relies on a graph neural network enhanced by attention-based components. By processing object features with geometric priors, the network is capable of performing multiple tasks, from linking coordinates into trajectories to inferring local and global dynamic properties. We demonstrate the flexibility and reliability of this approach by applying it to real and simulated data corresponding to a broad range of biological experiments.