Barbora Špačková, Henrik Klein Moberg, Joachim Fritzsche, Johan Tenghamn, Gustaf Sjösten, Hana Šípová-Jungová, David Albinsson, Quentin Lubart, Daniel van Leeuwen, Fredrik Westerlund, Daniel Midtvedt, Elin K. Esbjörner, Mikael Käll, Giovanni Volpe & Christoph Langhammer
Nature Methods 19, 751–758 (2022)
doi: 10.1038/s41592-022-01491-6
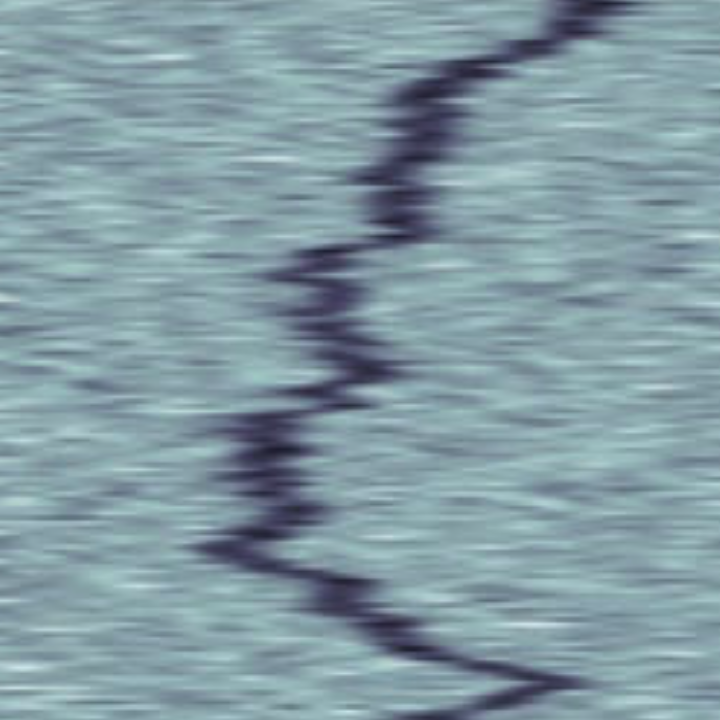
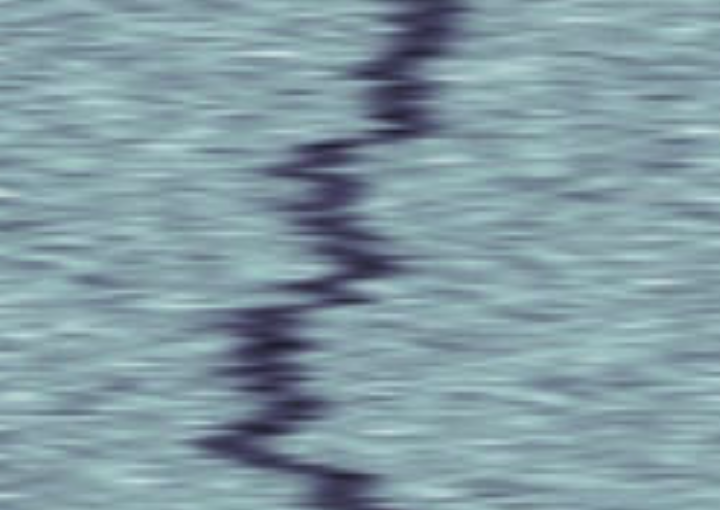
Label-free characterization of single biomolecules aims to complement fluorescence microscopy in situations where labeling compromises data interpretation, is technically challenging or even impossible. However, existing methods require the investigated species to bind to a surface to be visible, thereby leaving a large fraction of analytes undetected. Here, we present nanofluidic scattering microscopy (NSM), which overcomes these limitations by enabling label-free, real-time imaging of single biomolecules diffusing inside a nanofluidic channel. NSM facilitates accurate determination of molecular weight from the measured optical contrast and of the hydrodynamic radius from the measured diffusivity, from which information about the conformational state can be inferred. Furthermore, we demonstrate its applicability to the analysis of a complex biofluid, using conditioned cell culture medium containing extracellular vesicles as an example. We foresee the application of NSM to monitor conformational changes, aggregation and interactions of single biomolecules, and to analyze single-cell secretomes.