Daniel Midtvedt, Vasilii Mylnikov, Alexander Stilgoe, Mikael Käll, Halina Rubinsztein-Dunlop and Giovanni Volpe
Nanophotonics, 11(14), 3189-3214 (2022)
doi: 10.1515/nanoph-2022-0197
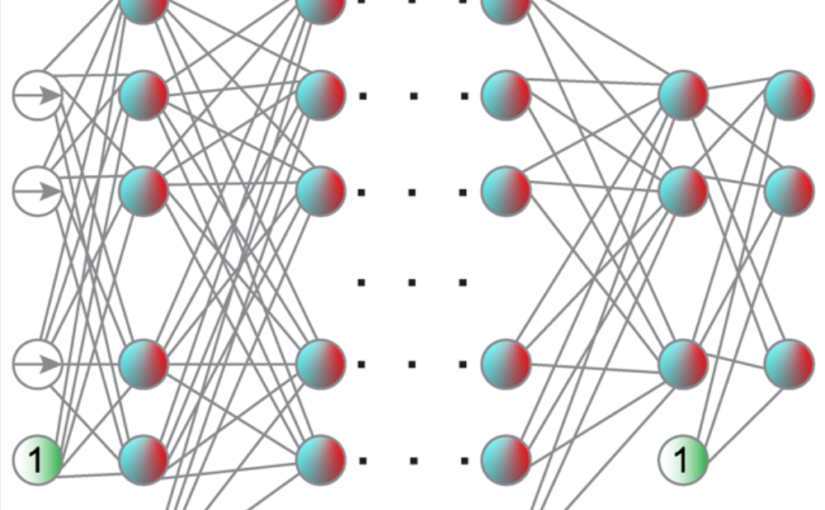
The deep-learning revolution is providing enticing new opportunities to manipulate and harness light at all scales. By building models of light–matter interactions from large experimental or simulated datasets, deep learning has already improved the design of nanophotonic devices and the acquisition and analysis of experimental data, even in situations where the underlying theory is not sufficiently established or too complex to be of practical use. Beyond these early success stories, deep learning also poses several challenges. Most importantly, deep learning works as a black box, making it difficult to understand and interpret its results and reliability, especially when training on incomplete datasets or dealing with data generated by adversarial approaches. Here, after an overview of how deep learning is currently employed in photonics, we discuss the emerging opportunities and challenges, shining light on how deep learning advances photonics.