Mite Mijalkov, Giovanni Volpe, Joana B Pereira
Cerebral Cortex, bhab237 (2021)
doi: 10.1093/cercor/bhab237
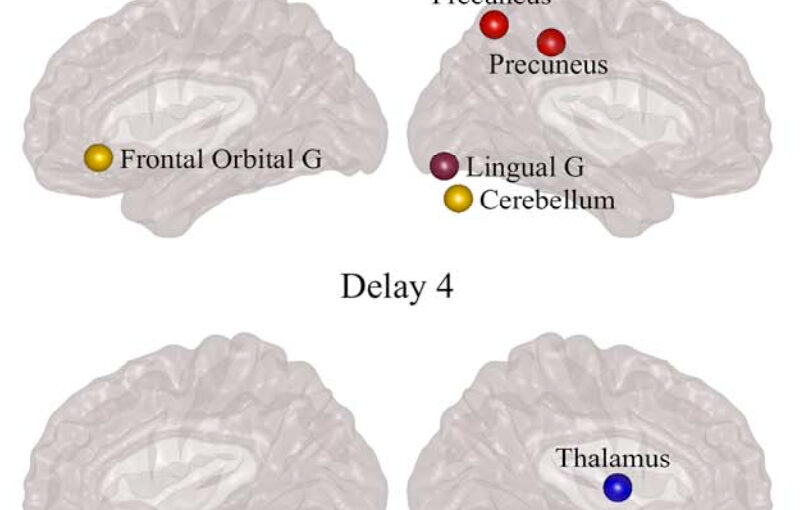
Parkinson’s disease (PD) is a neurodegenerative disorder characterized by topological abnormalities in large-scale functional brain networks, which are commonly analyzed using undirected correlations in the activation signals between brain regions. This approach assumes simultaneous activation of brain regions, despite previous evidence showing that brain activation entails causality, with signals being typically generated in one region and then propagated to other ones. To address this limitation, here, we developed a new method to assess whole-brain directed functional connectivity in participants with PD and healthy controls using antisymmetric delayed correlations, which capture better this underlying causality. Our results show that whole-brain directed connectivity, computed on functional magnetic resonance imaging data, identifies widespread differences in the functional networks of PD participants compared with controls, in contrast to undirected methods. These differences are characterized by increased global efficiency, clustering, and transitivity combined with lower modularity. Moreover, directed connectivity patterns in the precuneus, thalamus, and cerebellum were associated with motor, executive, and memory deficits in PD participants. Altogether, these findings suggest that directional brain connectivity is more sensitive to functional network differences occurring in PD compared with standard methods, opening new opportunities for brain connectivity analysis and development of new markers to track PD progression.